Summary
This tutorial describes a batch run of four single site analyses. Each analysis examines the performance of germplasm replicated three times in a balanced incomplete complete block design.
- Restore from Previous Tutorial
- Introduction
- Select Dataset to Analyze
- Load Data into Breeding View
- Run Analysis
- Review Results
- Quality Assurance
- Report
- Heritability Table
- Combined File of Predicted Means
- Individual Environment Reports
- Environment Report Summary
- Genotypes by Environment Sorted by BLUPs
- Summary of Traits by BLUP
- Genetic Correlations Between Traits
- Summary Statistics for Individual Trait Raw Data
- Estimated Heritability of Individual Trait
- Genotypes by Trait Sorted by BLUEs
- Standard Errors of Difference
- Wald/F Test
- Diagnostic Residual Plots of Individual Traits
- Upload BLUEs & Summary Stats to BMS
- References
- Related Materials
Restore from Previous Tutorial
Screenshots and activities in this tutorial build upon work preformed in previous tutorials.
- If you are not following the maize tutorials in sequence, restore the Maize Tutorial database (.sql) to the end of the previous tutorial, Trial Data Collection, to match database contents with current tutorial.
Restoration File?: Maize Database Trial with Observations (.sql)
Introduction
Breeding View’s single site analysis produces adjusted means, best linear unbiased estimators and best linear unbiased predictors (BLUEs and BLUPs) per genotype, as well as summary statistics to describe the data. The next tutorial, Maize Multi-Site Analysis, uses the summary statistics and adjusted means (BLUEs) to perform a genotype by environment (GxE) analysis. Adjusted means can also be used in a QTL (quantitative trait loci) analysis pipeline.
Select Dataset to Analyze
- Open Single Site Analysis from the Statistical Analysis menu of the Workbench. Select Browse to find the 4 Location Trial dataset.
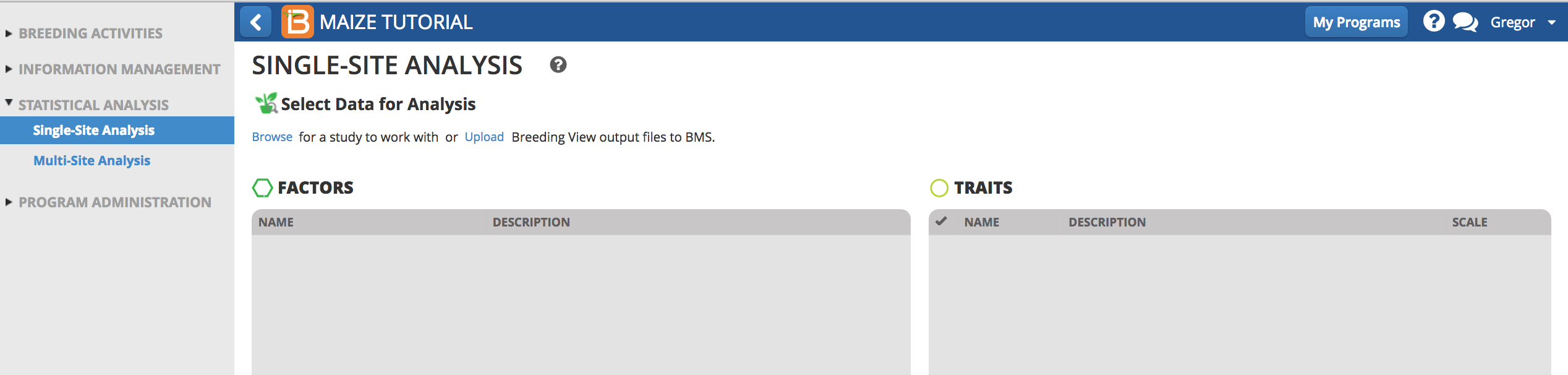
- Select 2017 Performance Trial.
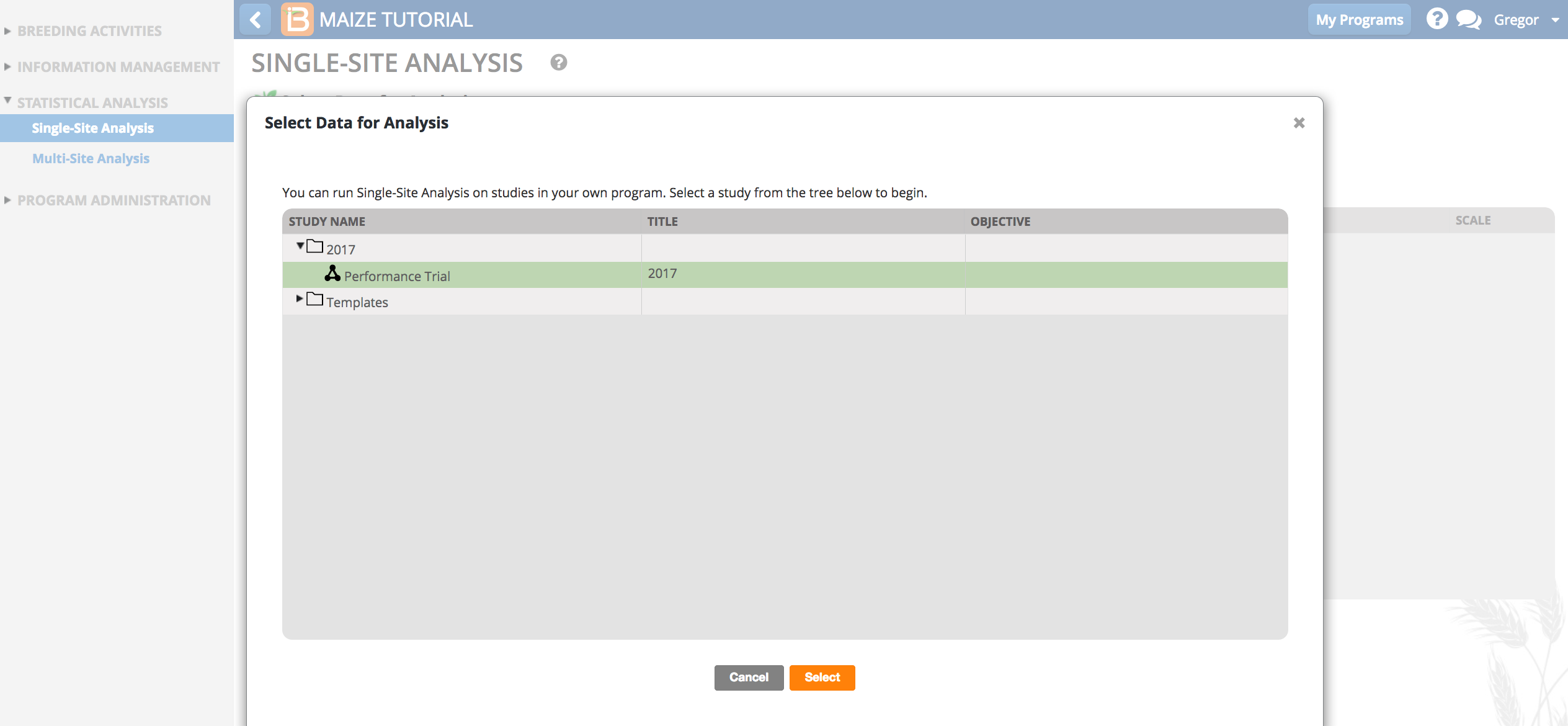
- Review the factors and make sure the six phenotypic traits measured in this trial dataset are selected, and click Next.
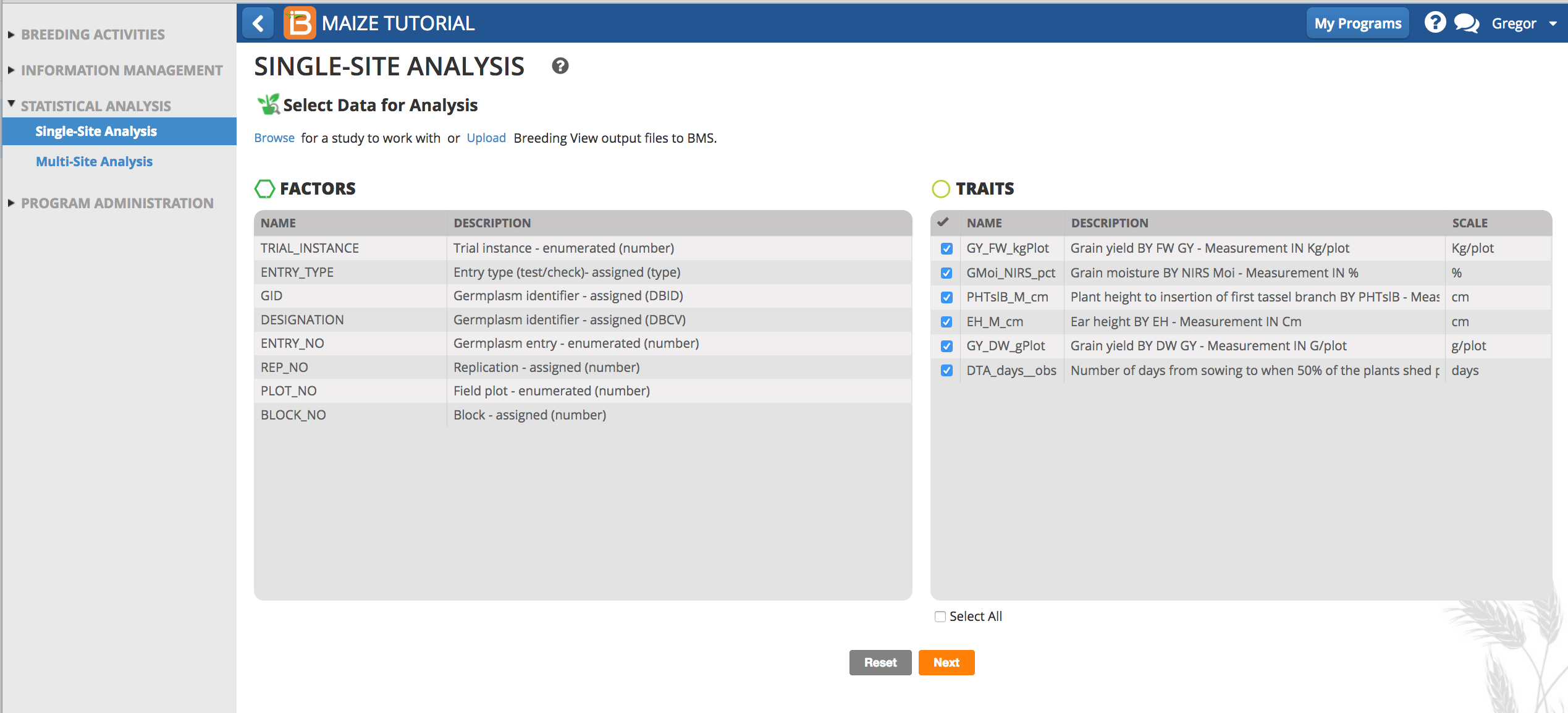
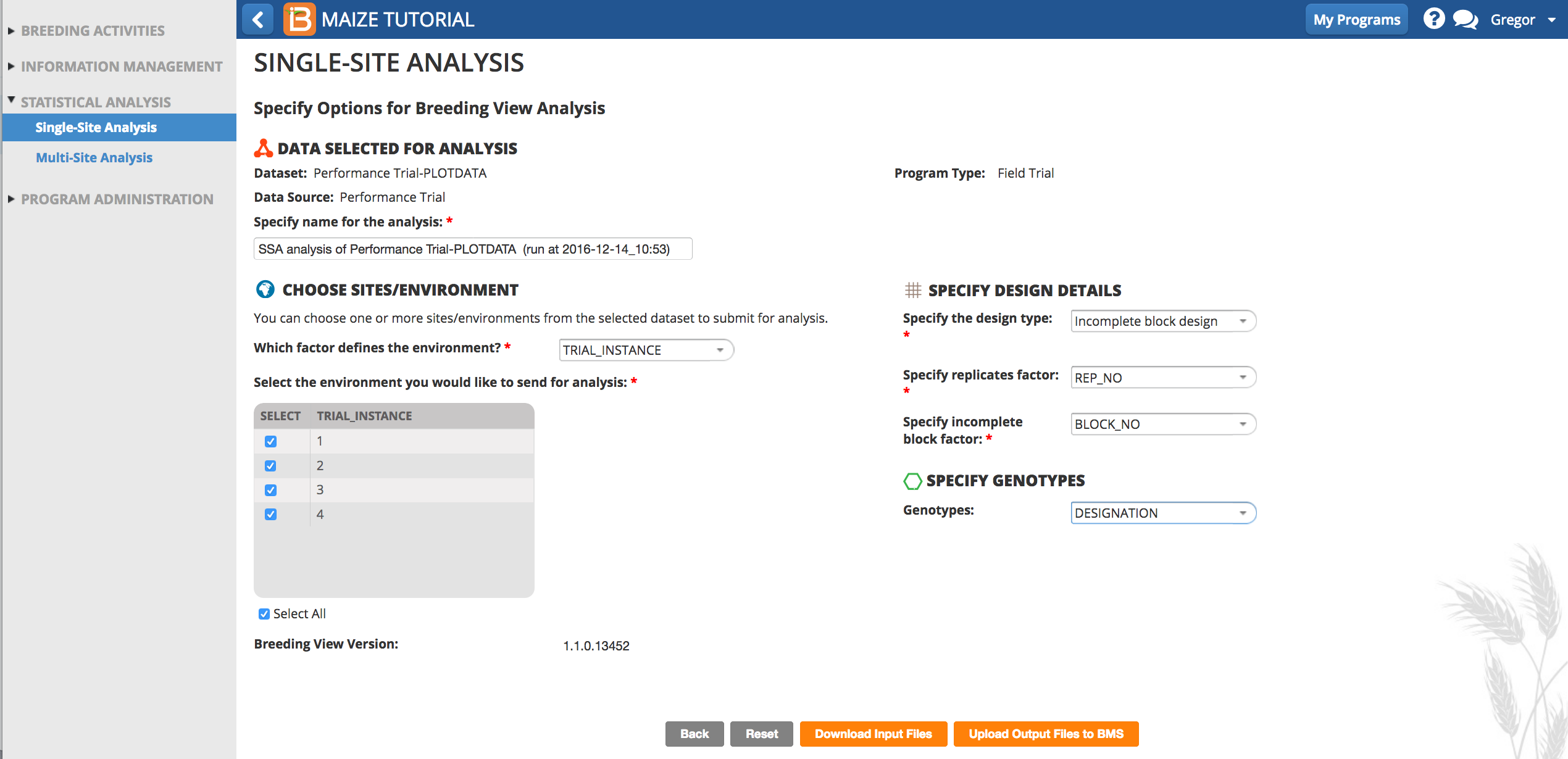
- Specify Analysis Conditions
- Use the default analysis name.
- Select Incomplete Block Design for the design type.
- The factor that defines environment is TRIAL_INSTANCE.
- Select all 4 instances to perform 4 individual single site analyses
- REP_NO is the factor in this dataset that defines replications.
- DESIGNATION defines the germplasm factor to be used in the analysis.
- Select Download Input Files.
- Decompress, or unzip, the input folder to reveal the two files for input into the Breeding View application.
Load Data into Breeding View
BMS v4.0 is a server application. Breeding View (BV) is a Windows compatible desktop application (see Install Breeding View). Trial data exported from the BMS are perfectly formatted for analysis by the BV application.
- Launch the Breeding View application on your local computer and select Open Project.
- Load trial details into the Breeding View application. Browse to the .xml file and open.

- Select Add to Project.
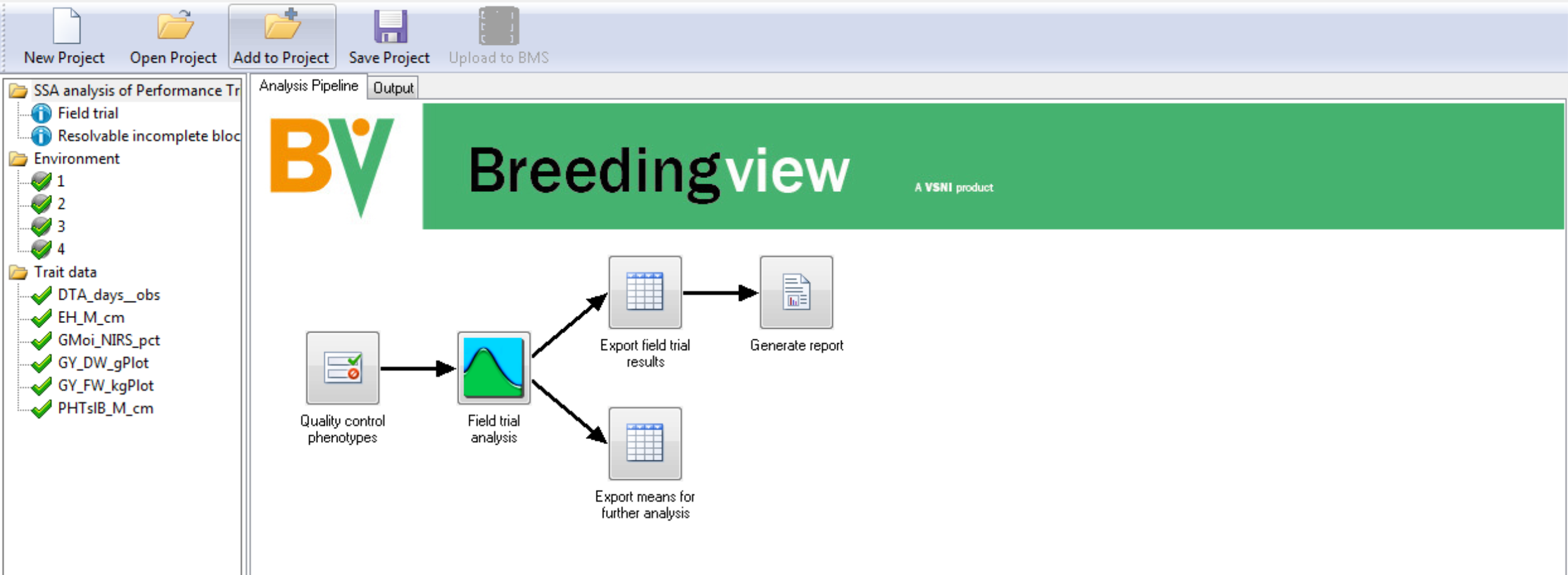
Run Analysis
The analysis pipeline includes a set of connected nodes, which can be used to run and configure pipelines.
- Right click on the Quality Control Phenotypes. Run the analysis pipelines for the selected 4 environments.
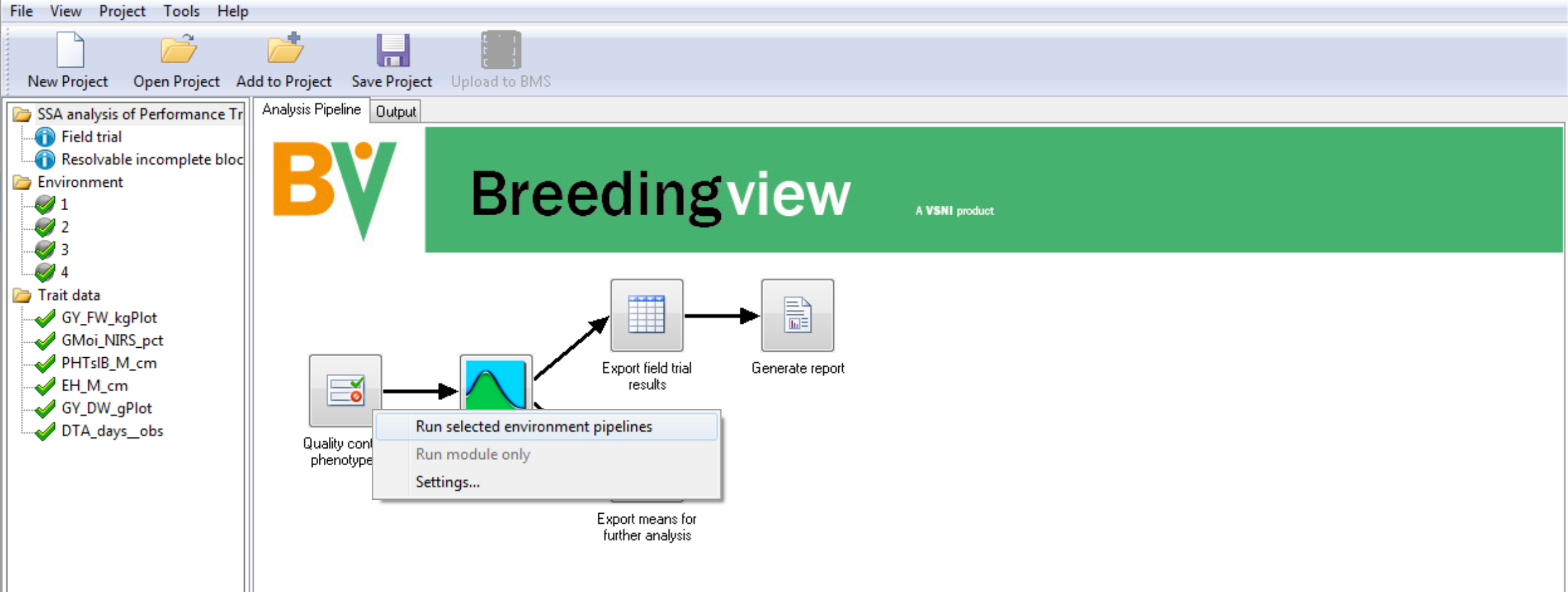
When the analysis is complete a popup notifies the user.
- Select OK to view the analysis results.
All of the nodes in the analysis pipeline are green when the analyses are complete.
- Select the Quality Assurance tab.
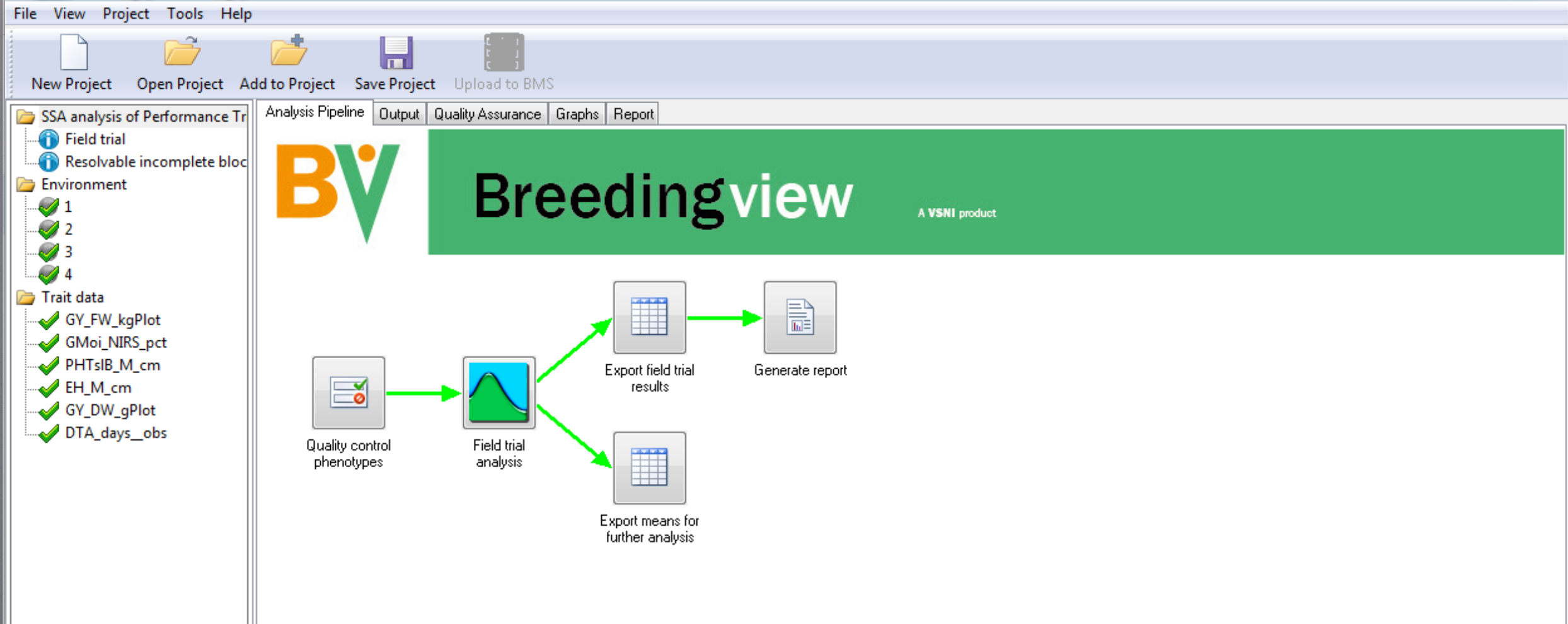
Review Results
Quality Assurance
Breeding View provides an overview of potentially influential measurements to help users identify and possibly exclude observations. Influential observations may reflect true genotypic variation and care should be taken not to exclude these data from the trial. Observations that deserve exclusion are obvious errors or measurements influenced by heterogeneous environmental variation within a block, like damage to a single plot.
- Select and open the report of the trial conducted at Environment 2.
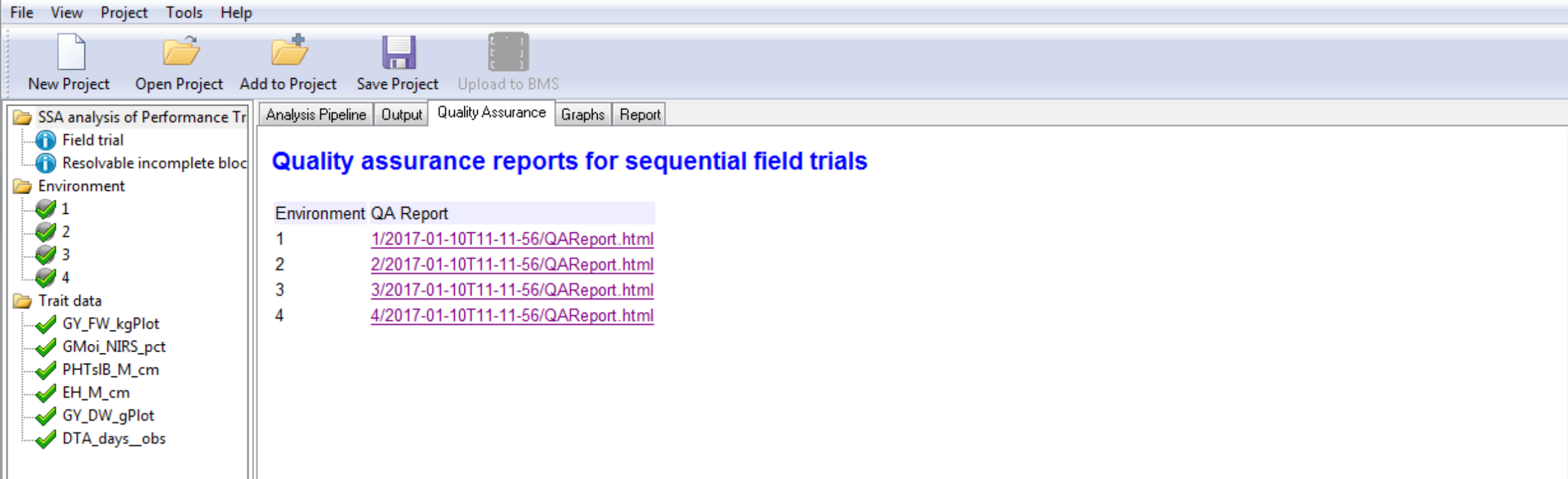
The table displays potential influential observations identified by the raw data method, which identifies observations exceeding 1.5 times the interquartile range, and the residual method, which identified standardized residuals by mixed model analysis.
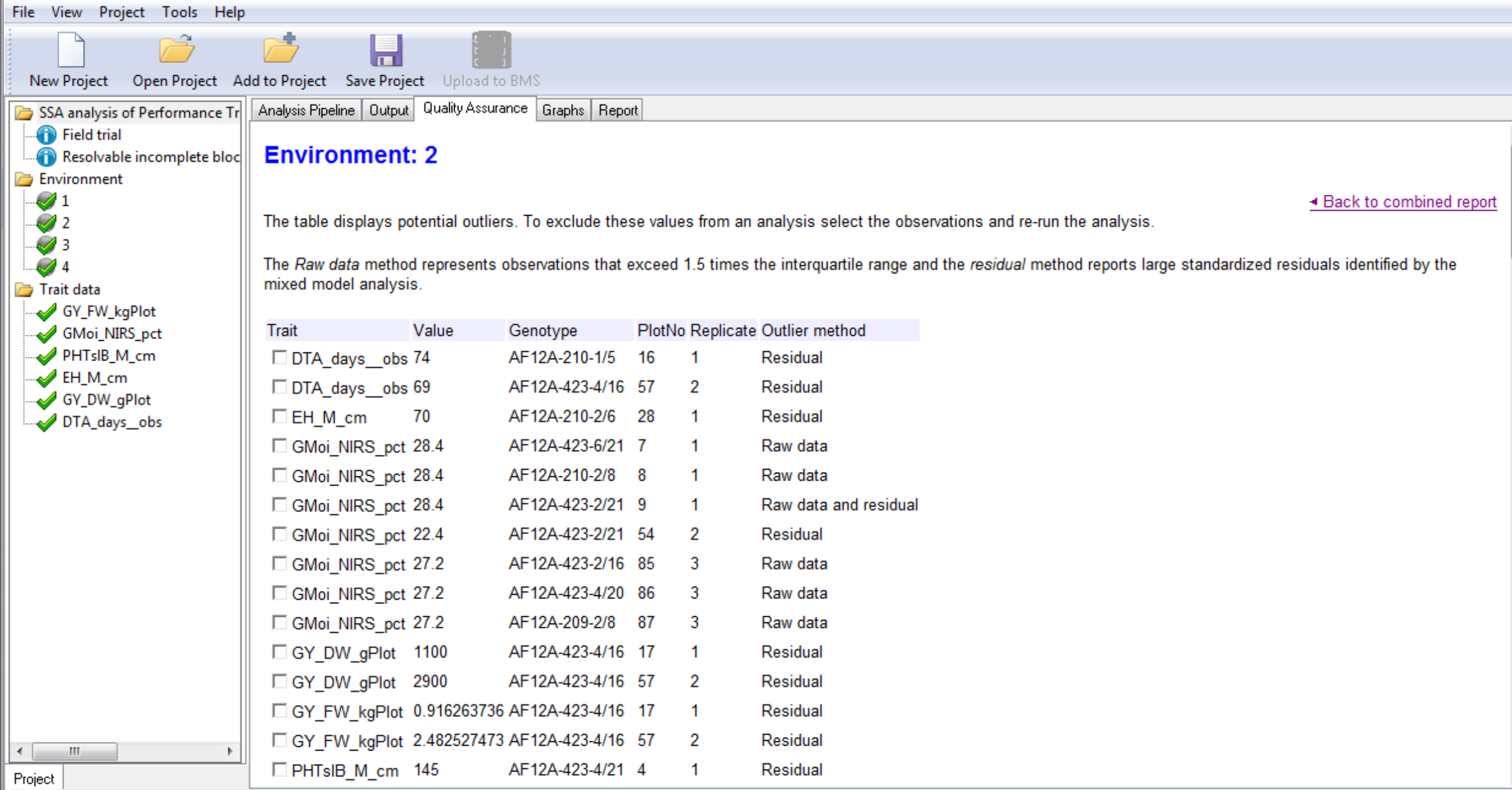
Box plots are provided to graphically illustrate influential measurements.
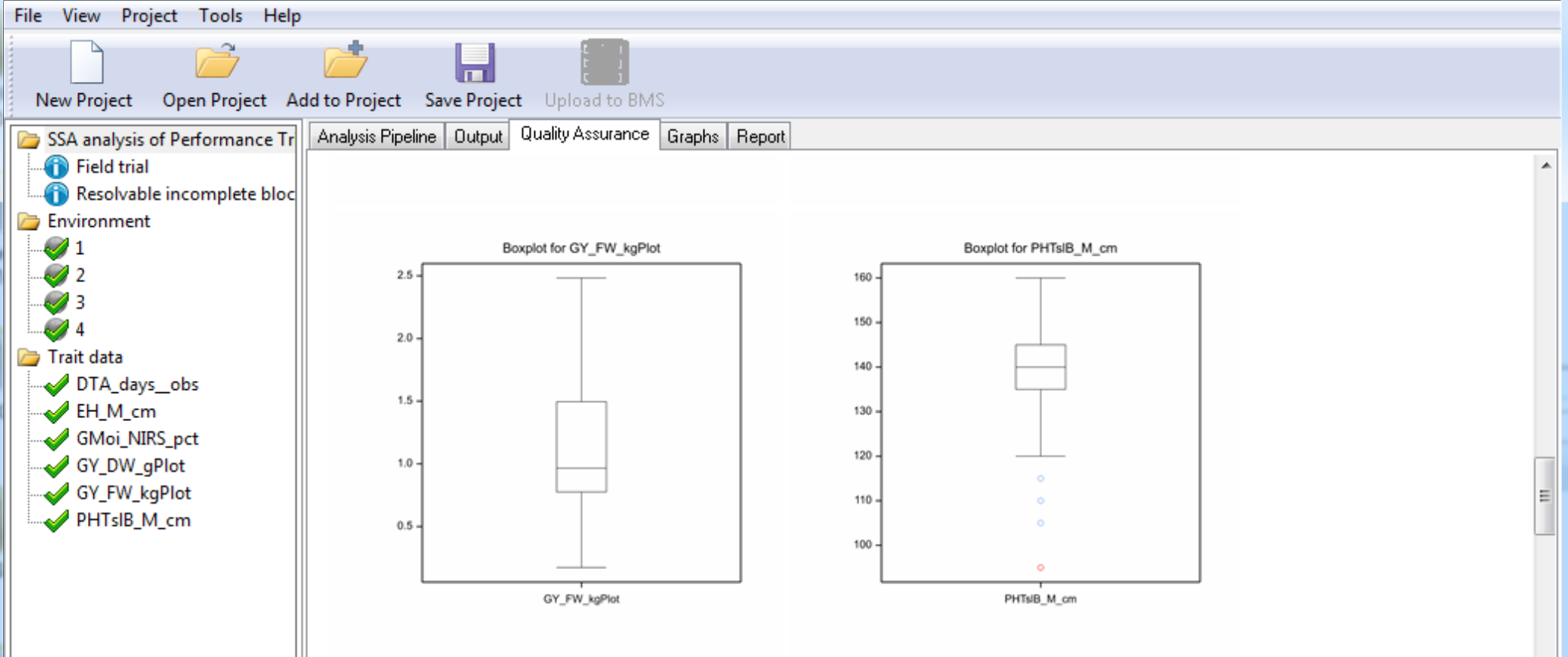
Notice that plant height (PHTsIB_M_cm) has a potential outlier measurement highlighted in red.
Report
This section of the tutorial provides a brief guide to interpretation of the results, including graphs, included under the Report tab.
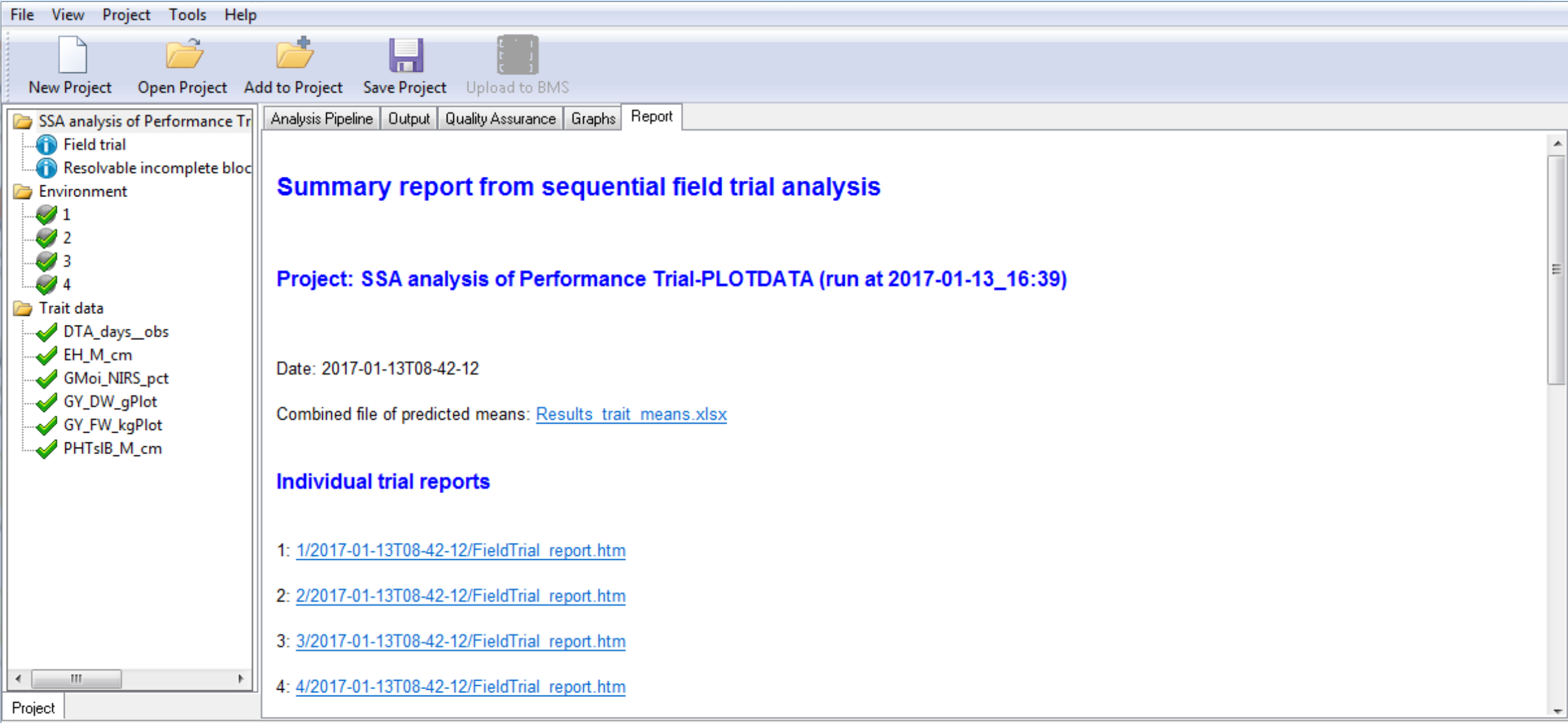
Report Tab Contents
- Heritability Table
- Combined File of Predicted Means: Excel file of BLUEs and BLUPs
- Links to Individual Environment Reports
Heritability Table
The heritability table summarizes the generalized heritabilities calculated for each trait by location as described by Oakey et al. 2006. The method uses the average pairwise prediction error variance to obtain genetic and error covariance matrices and allows for the estimation of heritability in unbalanced data with complex error and genetic structures. If a model cannot be fitted to the trait data, such as when there is no variability in the trait measurement, that trait will not be included in the heritability table.
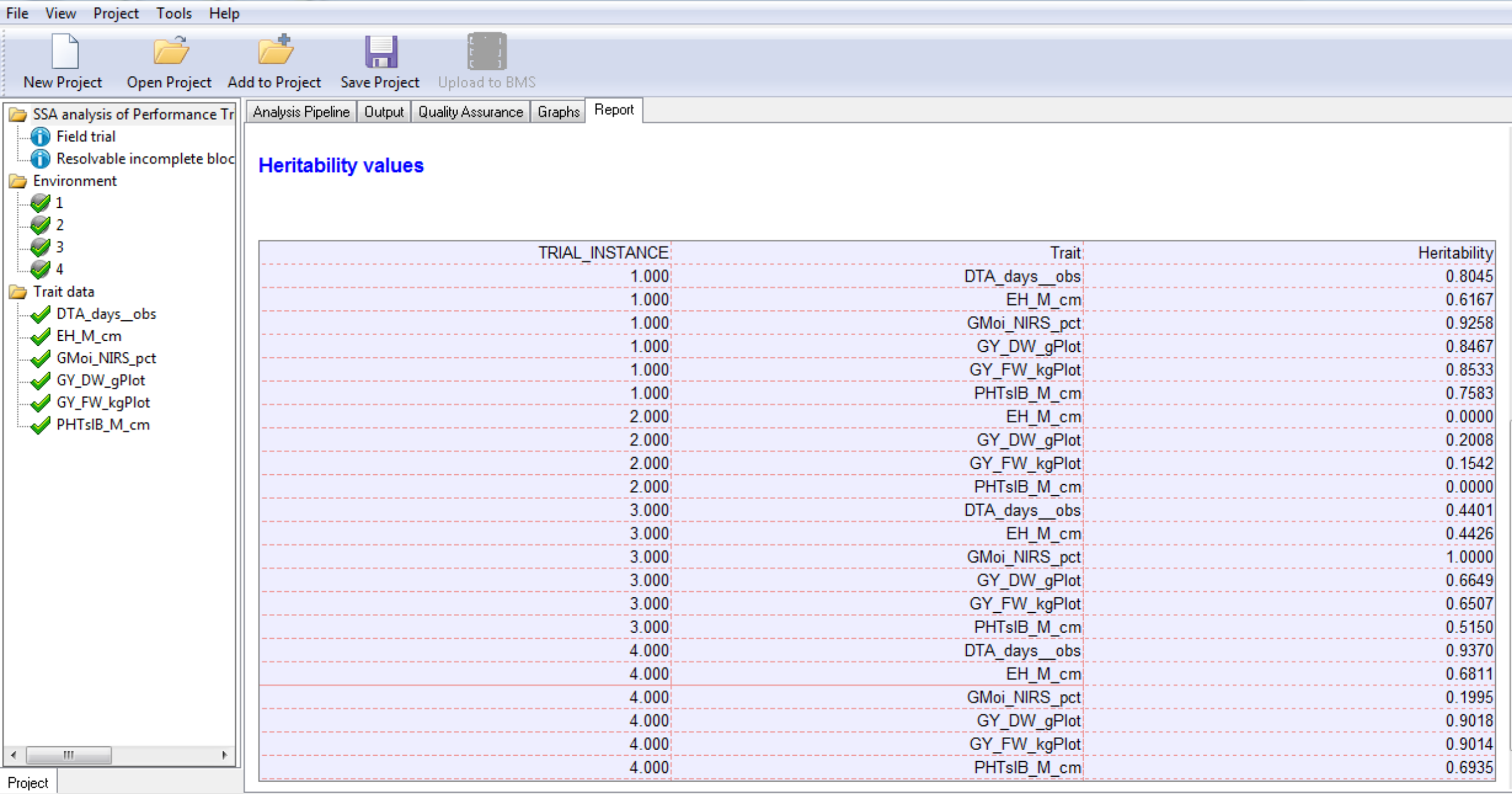
Combined File of Predicted Means
- Select the link to the combined file of predicted means and open in Excel. Non-informative traits that could not be fitted with a model will appear as missing data.
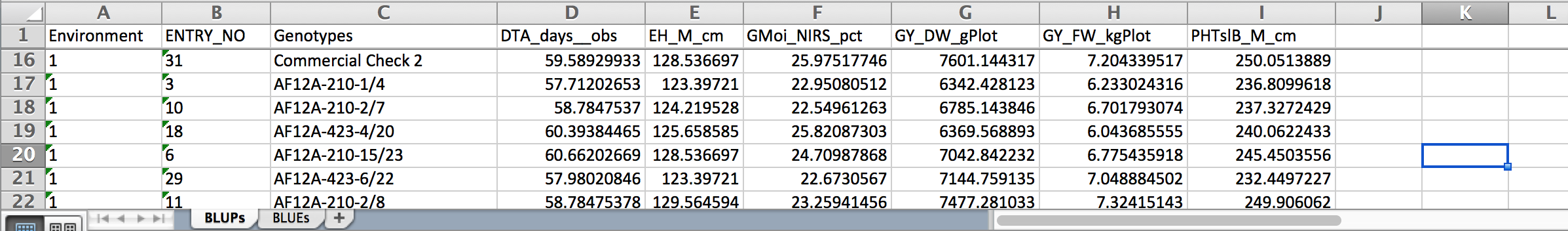
BLUPs across all locations
Individual Environment Reports
Although the four locations are summarized together in the heritability table, each location is represented by an individual analysis. Select the link to the Environment 4 individual trial report to review the analysis performed at this location.
Individual Environment Reports Include:
- File of predicted means: Link to Excel (.xls) file containing an environment-specific subset of BLUEs and BLUPs
- Best Genotypes Table: Best genotypes as defined by BLUPs sorted by factors defined in the report options
- Summary of Traits: A table presenting the minimum, mean, maximum, and heritability for each trait within this location based on BLUPs
- Estimated Genetic Correlations Between Traits
- Principle Components Biplot
- Individual Trait Analyses: Summary statistics of raw data, heritability, sorted genotype table (BLUEs), standard errors of differences, and residual diagnostic plots.?
Environment Report Summary
The environment report provides the project name, the environment name, the field design, along with a date the stamp for the analysis. Users are presented a link to the adjusted means data for this environment, which is a subset of the data presented in the combined mean file reporting all locations. Users are also notified about analysis failures.
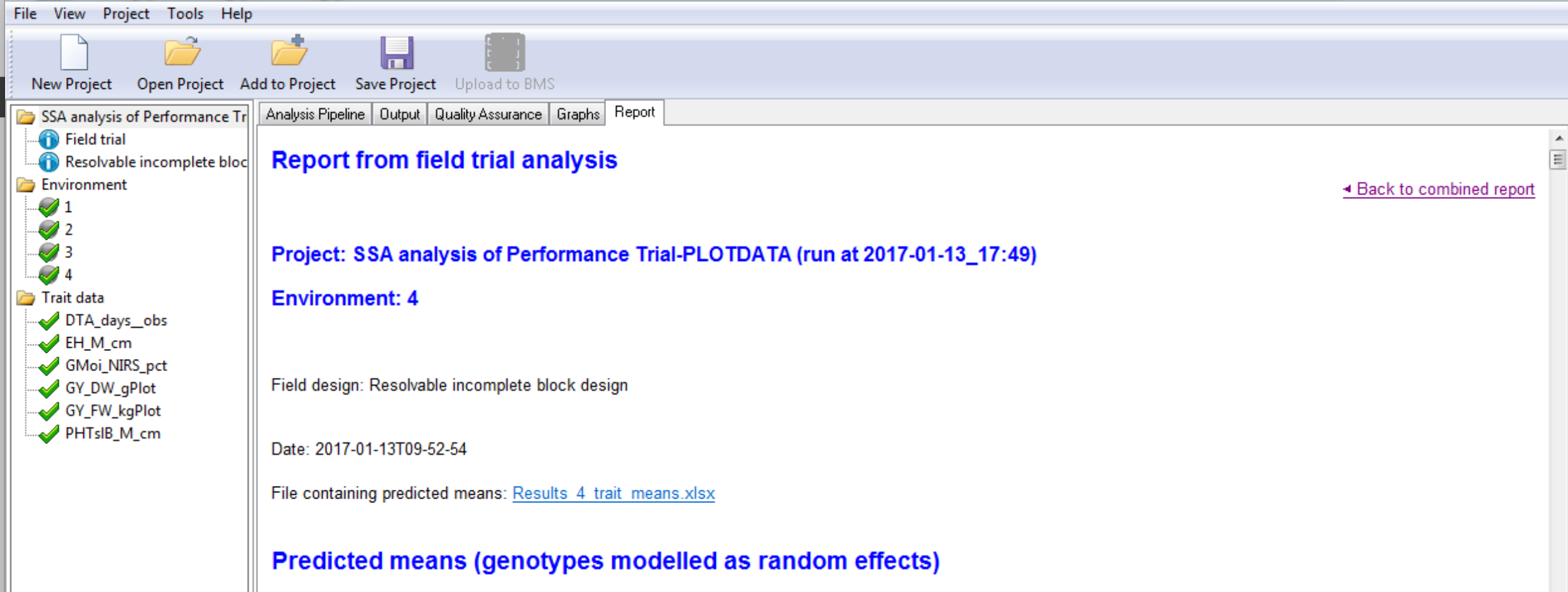
Genotypes by Environment Sorted by BLUPs
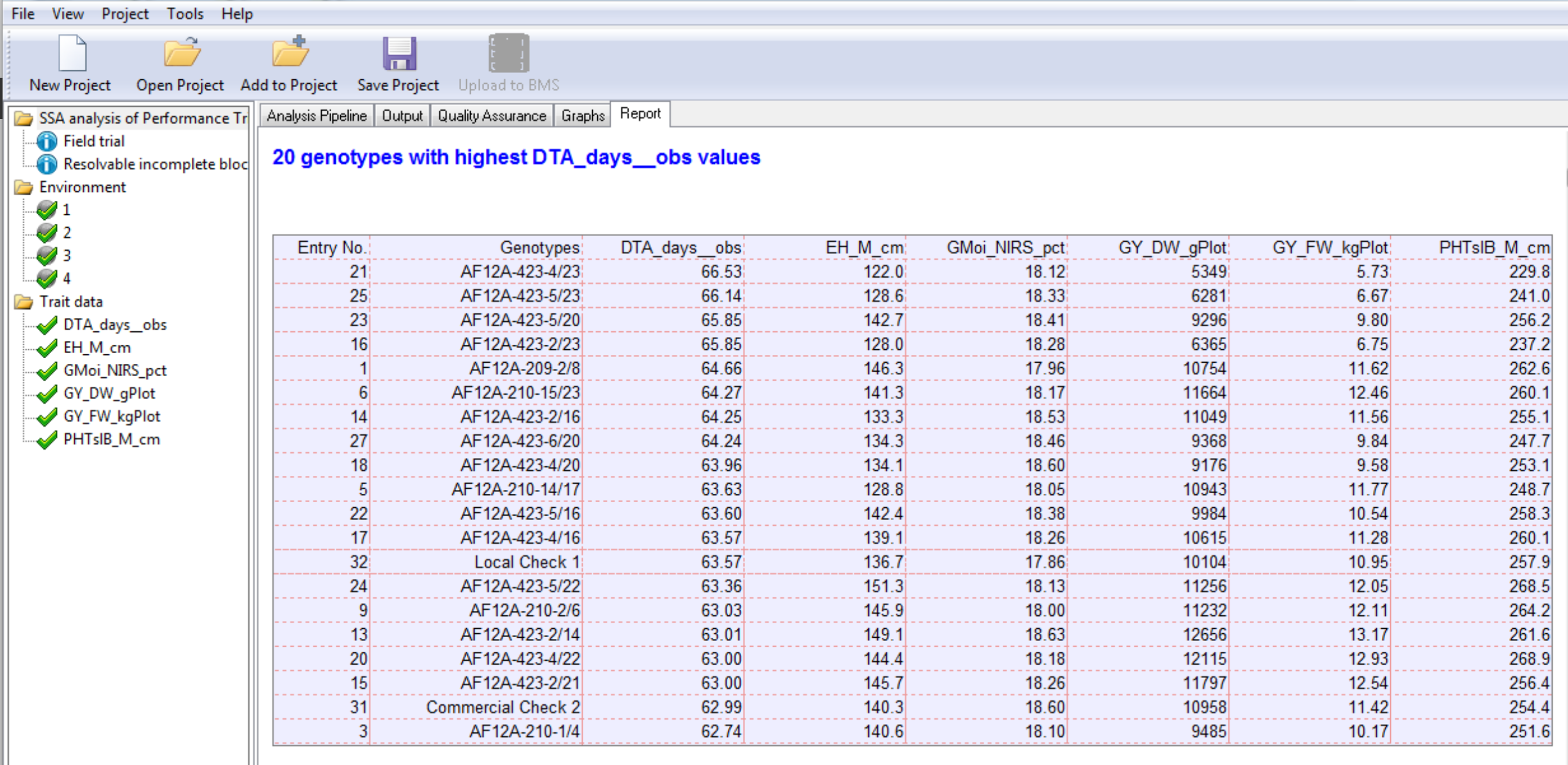 20 Best Genotypes at Environment 4 Sorted by Days to Anthesis: This table is the 20 best genotypes sorted by BLUPs for days to anthesis in descending order, as defined by the settings under the Generate Report node. Note that while breeders generally select for high phenotypic values, days to anthesis is an exception - delayed maturity is generally selected against.
20 Best Genotypes at Environment 4 Sorted by Days to Anthesis: This table is the 20 best genotypes sorted by BLUPs for days to anthesis in descending order, as defined by the settings under the Generate Report node. Note that while breeders generally select for high phenotypic values, days to anthesis is an exception - delayed maturity is generally selected against. Summary of Traits by BLUP
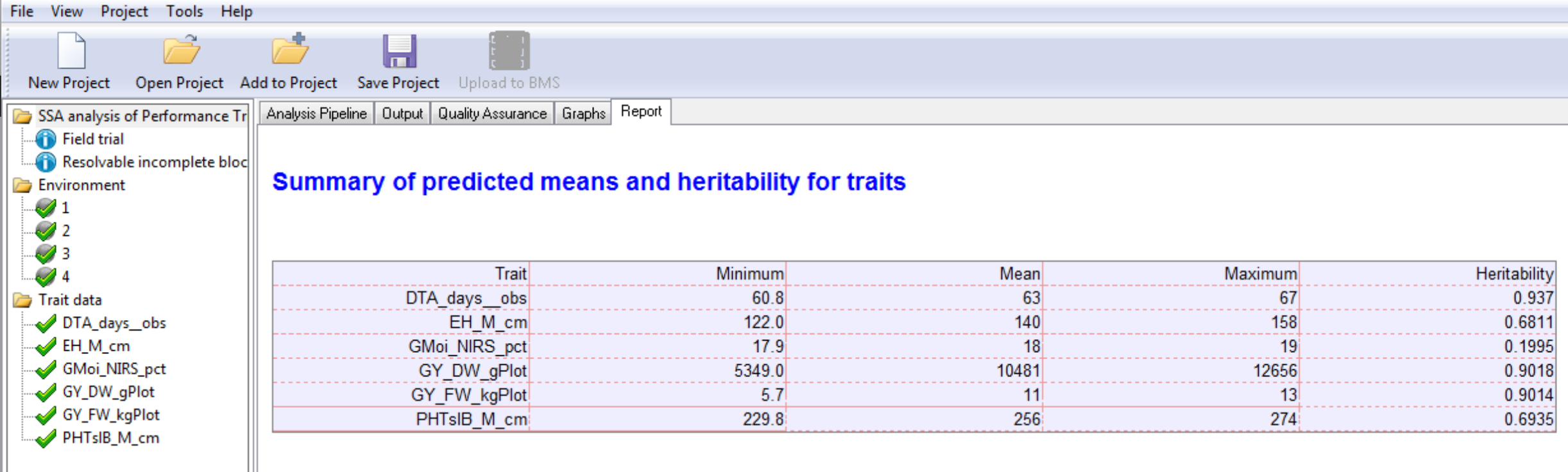
Environment 4 Summary of Traits: The minimum, mean, maximum, and heritability for each trait based on BLUPs
Genetic Correlations Between Traits
BLUPs Principle Components Biplot
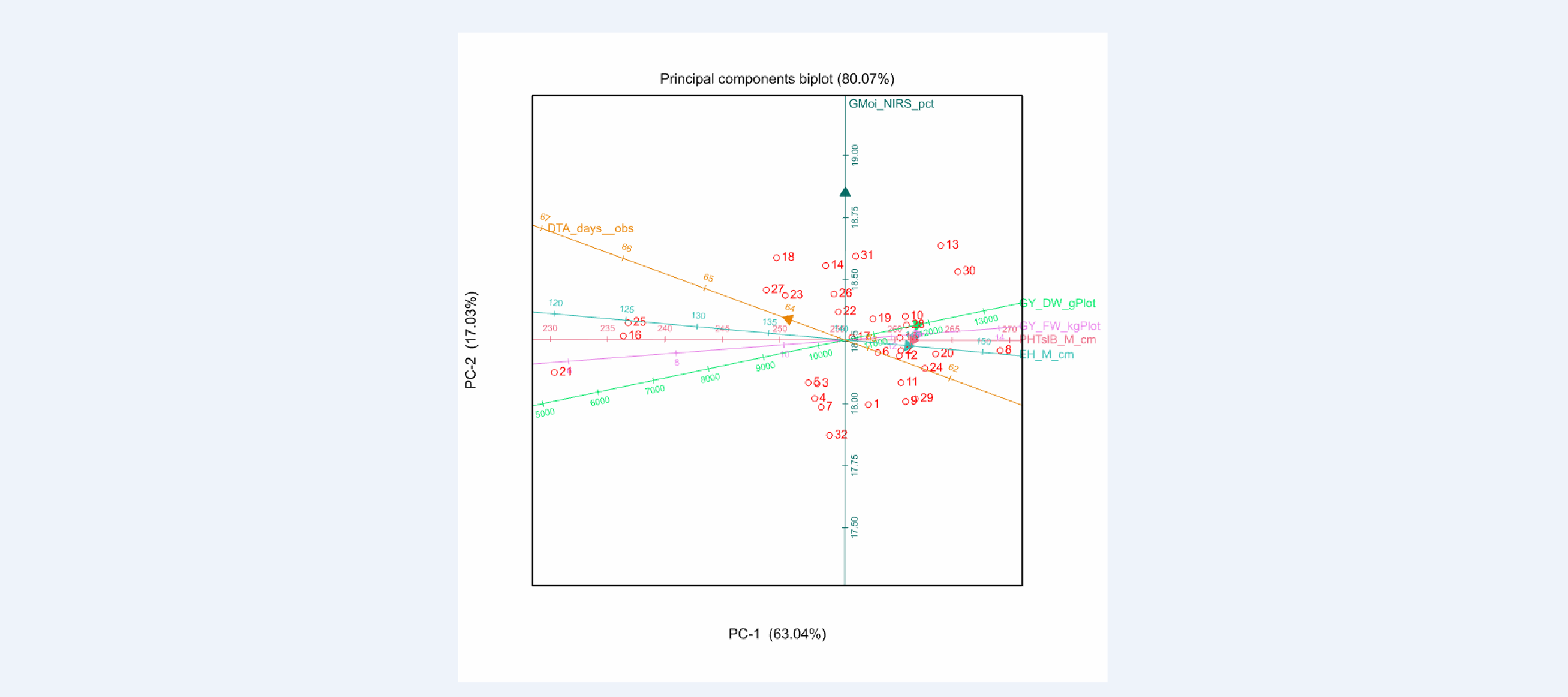
Environment 4 Principle Components Biplot of BLUPs:The biplot shows that field weight, grain yield dry & fresh weight (GY_DW_gPlot & GY_FW_kgPlot), plant height (PHTsIB_M_cm) , and ear height (EH_M_cm) are all positively correlated among each other (acute angle of vectors), and negatively correlated with anthesis date (DTA_days_obs) (obtuse angle). In addition, all traits are weakly correlated with grain moisture (GMoi_NIRS_pct) (right angle). These relationships can also be seen in the genetic correlation matrix below.
Genetic Correlation Matrix
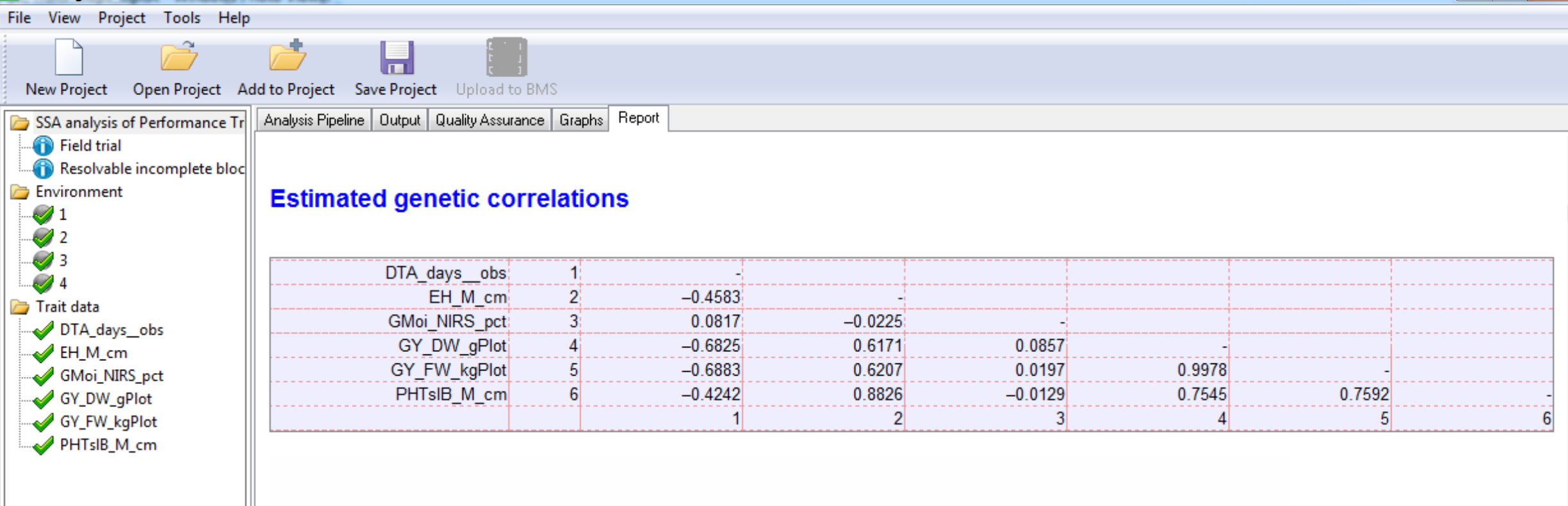
Environment 4 Estimated Genetic Correlations: Pairwise correlation (r) of phenotypic traits. There is a strong positive correlation (0.9978) between the related yield measures, grain yield and field weight (GY_DW_gPlot & GY_FW_kgPlot). These two yield traits are moderately negatively correlated (-0.6890 & -0.6830 respectively) to anthesis date (DTA_days_obs). In other words, late anthesis is correlated to low yield.
Summary Statistics for Individual Trait Raw Data
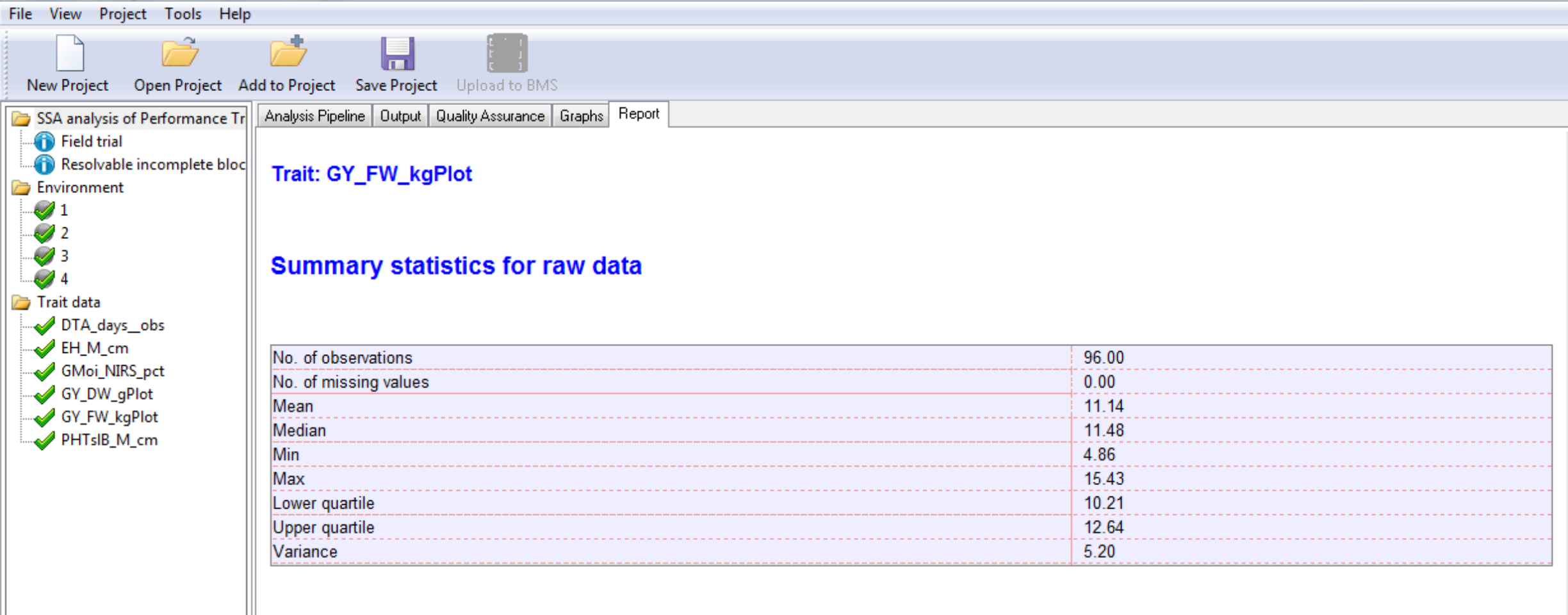
Environment 4 Summary Statistics for Grain Yield (GY_FW_kgPlot) based on raw data
Estimated Heritability of Individual Trait
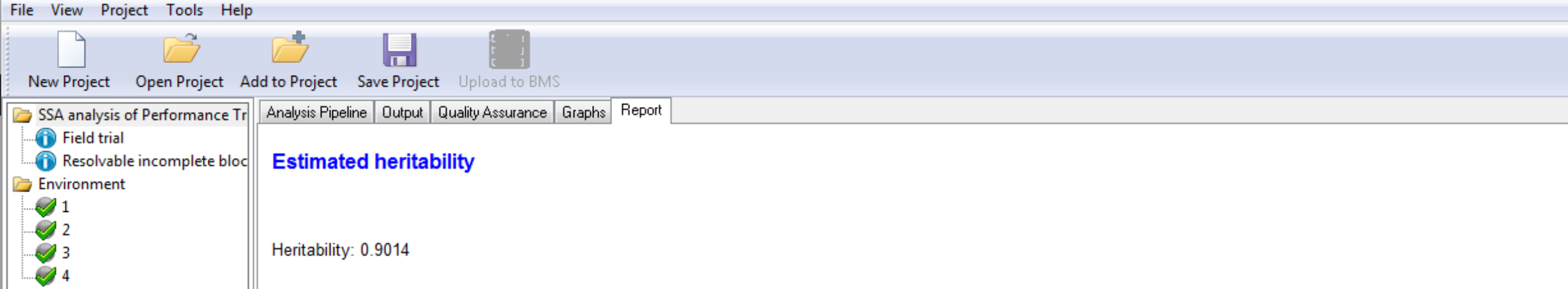 Estimated Heritability of Grain Yield (GY_FW_kgPlot) calculated at Environment 4
Estimated Heritability of Grain Yield (GY_FW_kgPlot) calculated at Environment 4Genotypes by Trait Sorted by BLUEs
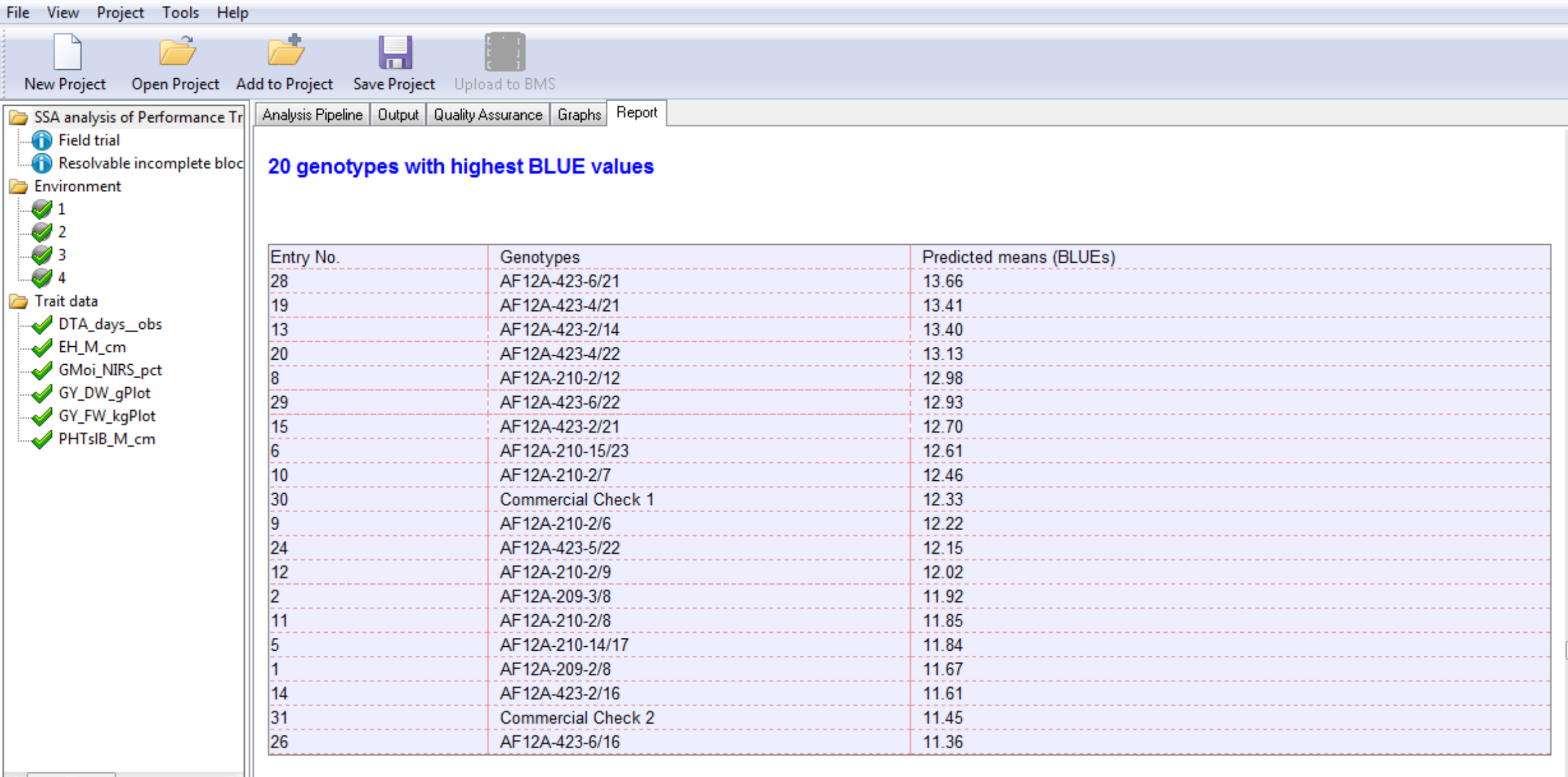
20 Best Grain Yield (GY_FW_kgPlot) Genotypes Calculated at Environment 4: Genotypes are sorted by BLUEs in descending order by value as specified in the Report Options.
Standard Errors of Difference
Two genotypes are considered different when their means are 2 times the standard error of the difference (SED), equivalent to LSD (Least Squared Difference). In general a breeder would use the average, and consider the minimum and maximum to have some sense of the differences in precision of comparisons among means.
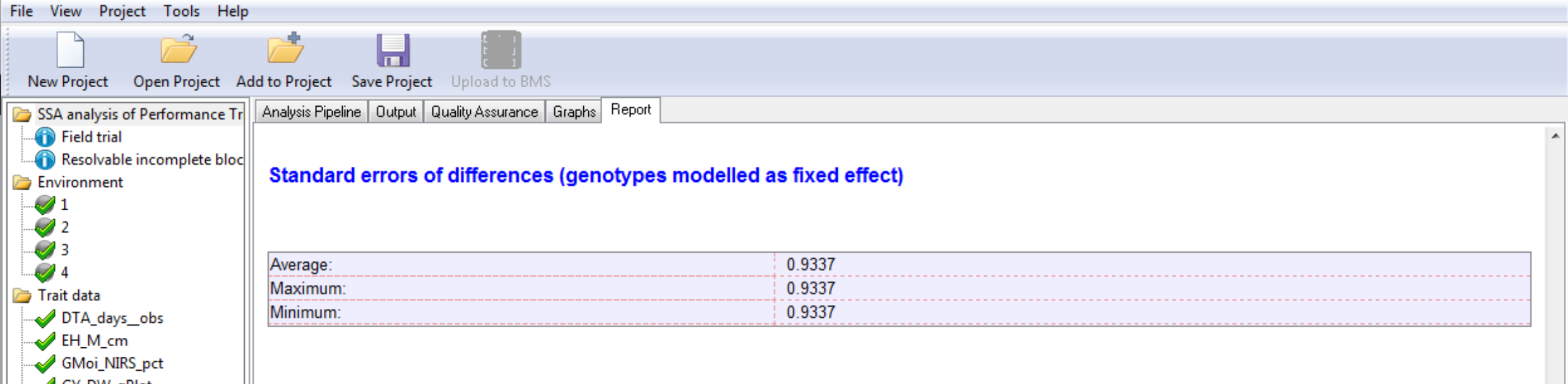
Standard Errors of Difference for Grain Yield (GY_FW_kgPlot): In a balanced design without missing data, like in this example the average, maximum, and minimum SED are equivalent.
Wald/F Test
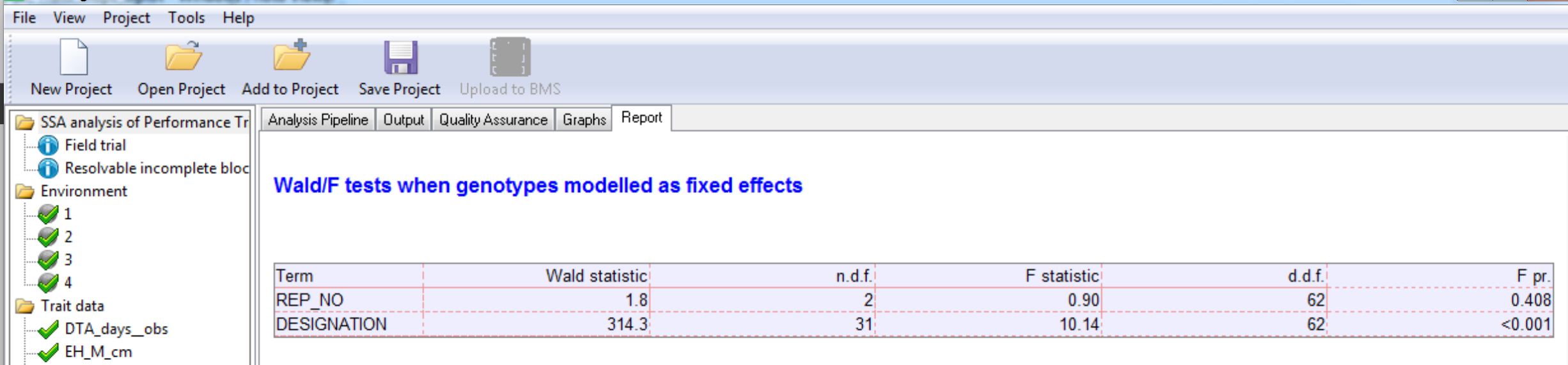
Diagnostic Residual Plots of Individual Traits
Diagnostic residual plots are used to check the model assumptions. Residuals are defined as the difference between the observed and fitted values. A good model “fit” for adjusted means will have residuals that should be independent and follow a normal distribution with a mean of zero and a constant variance. All but the independence assumption can be checked with the residual plots - independence follows from the randomization of the experimental design.
Utility of Each Diagnostic Plot
- Histogram of Residuals: Check for a normal, or Gaussian distribution, as well as centering on a mean of zero.
- Fitted-Value Plot: Check for constant variance as well as centering on zero. A random distribution, or “shot-gun pattern”, reflects constant variance. Positive or negative correlations between residuals and fitted values, or ‘loud-speaker-shaped’ distributions, point to violation point to a violation of the constant variance assumption
- Normal Plot: Check normality. Distribution in a straight line across the diagonal reflects a normal distribution. The Normal Plot has the same use as the Histogram of Residuals, but is generally a better visualization.
- Half-Normal Plot: Check normality. Distribution in a straight line across the diagonal reflects a normal distribution. The Half-Normal plot is the same as the normal plot, but considers the absolute value of residuals. This plot is useful with small data sets.
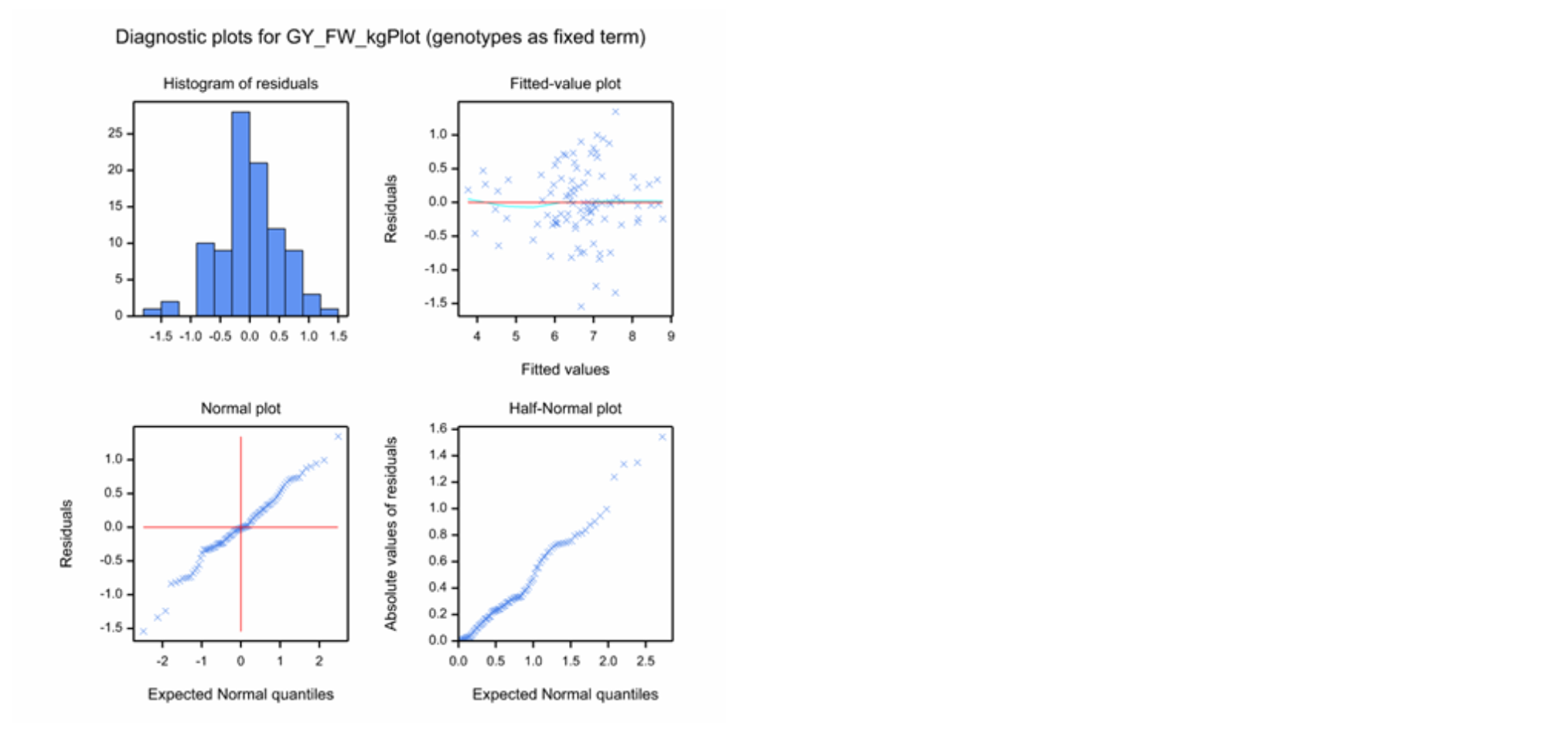
Diagnostic plots for Grain Yield (GY_FW_kgPlot) in Environment 4: Example of a continuous variable exhibiting a good model fit
Upload BLUEs & Summary Stats to BMS
- Return to the Breeding Management System Single-Site Analysis. Select Upload Output Files to BMS to save the adjusted means and summary statistics to the BMS database for use in a subsequent genotype by environment (GxE) analysis.
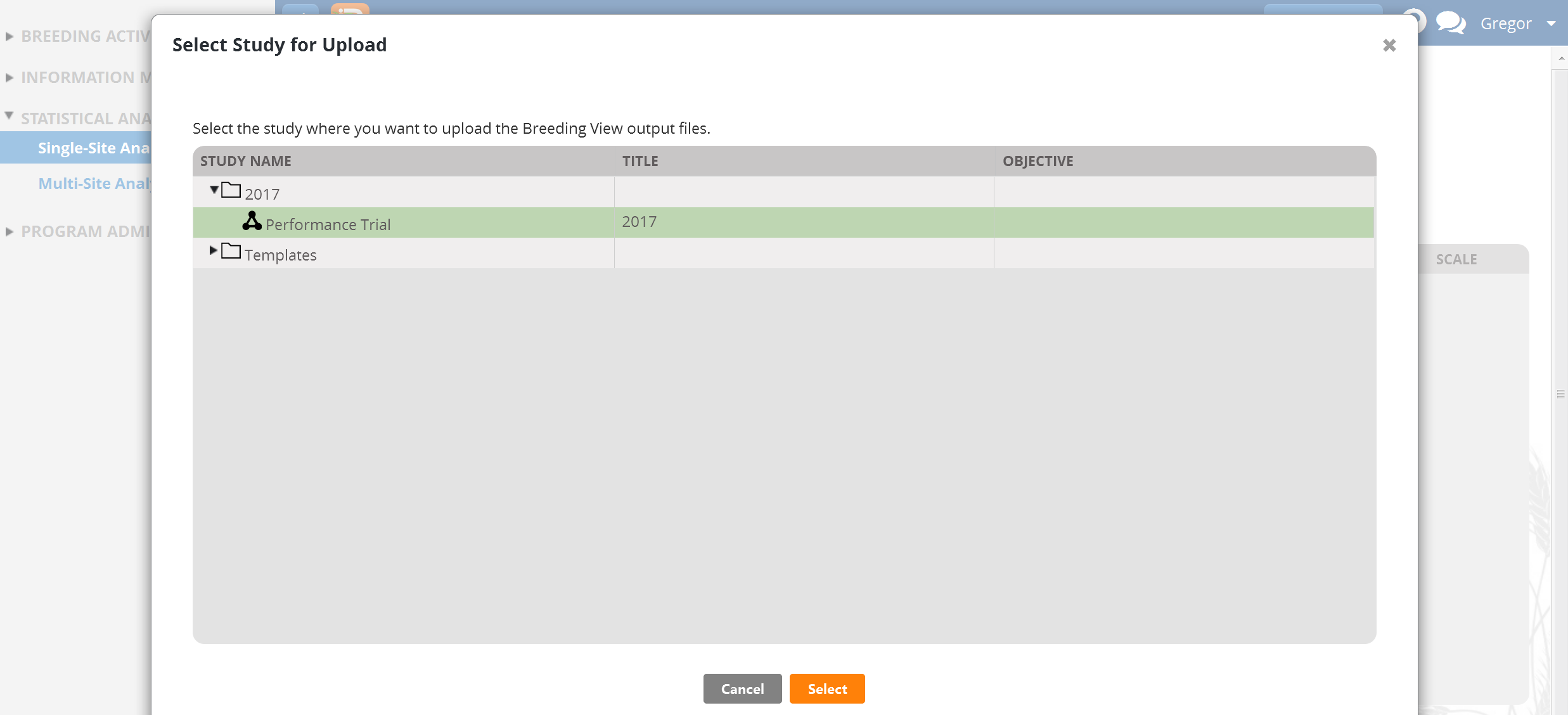
- Highlight the 2017 Performance Trial and Select.
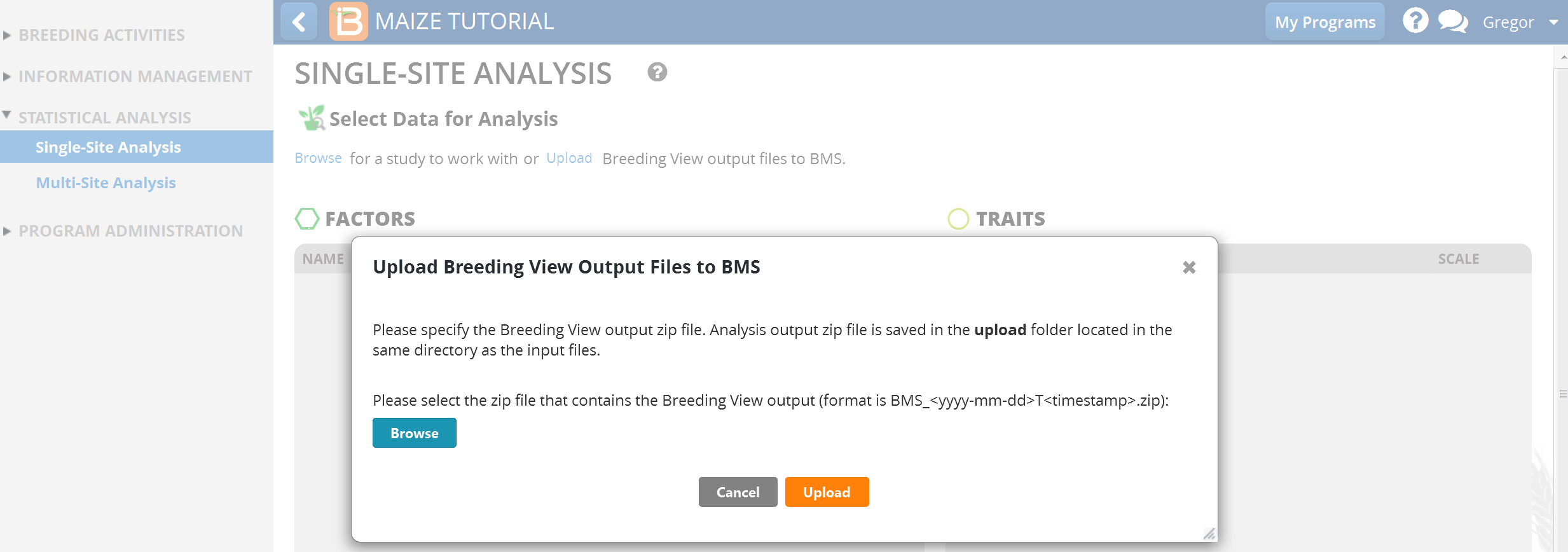
- Select Browse.
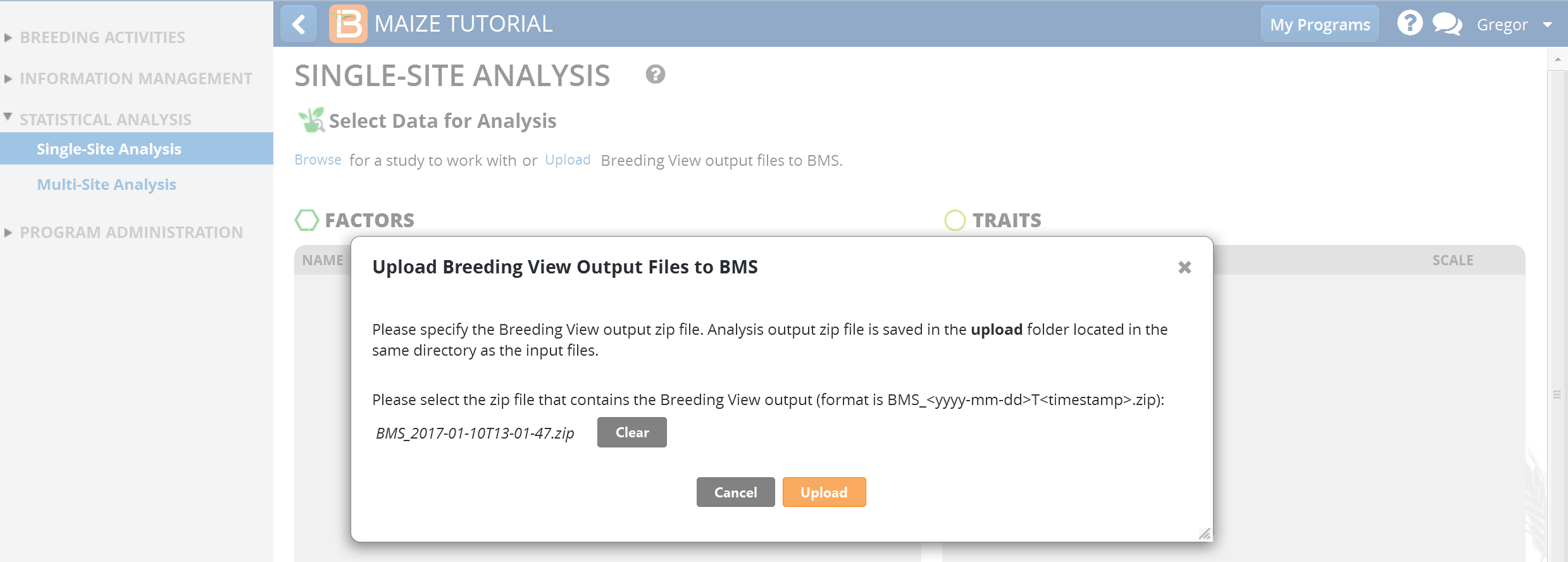
- Browse to the zipped Breeding View Upload file. The zip file is date and time stamped, and can be found within the upload folder. Select and upload.
Once the import is successful, the means and summary statistics from the single site analysis are available to perform a genotype by environment analysis.
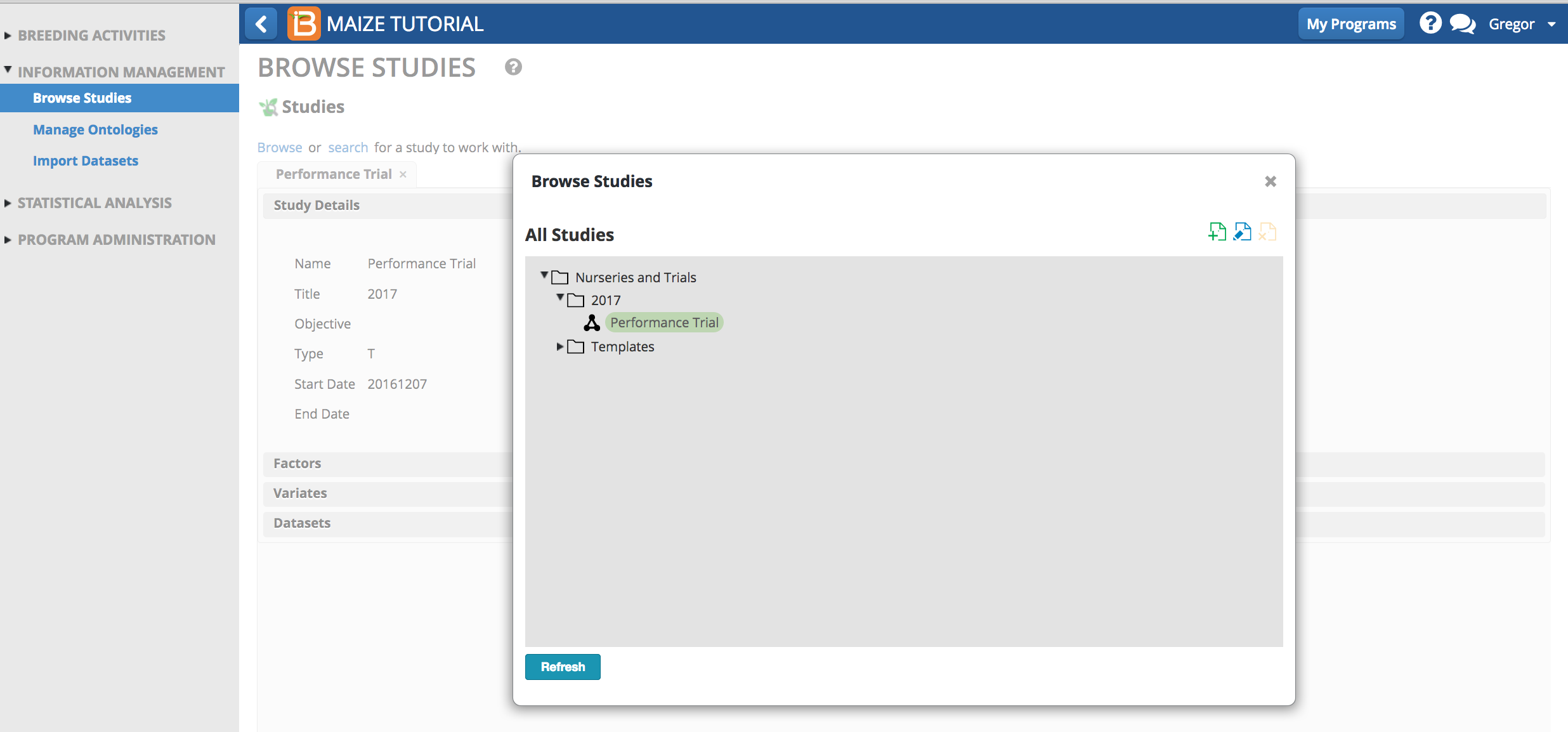
- Confirm upload by selecting the Performance Trial from the Browse Studies menu option.
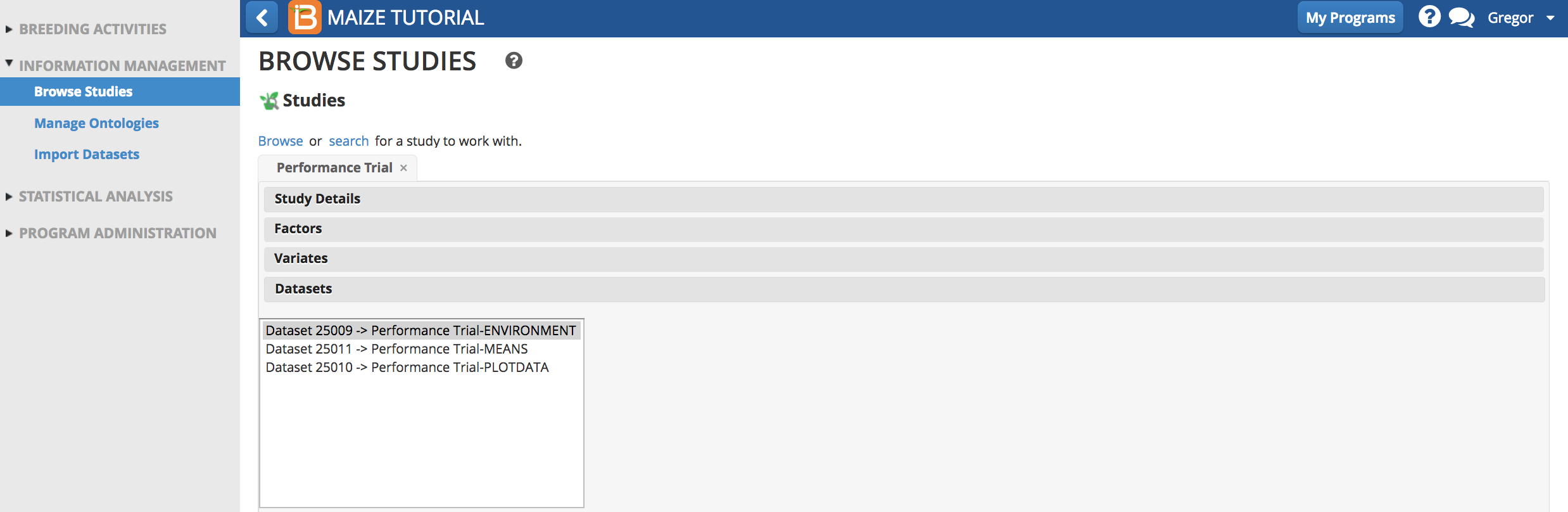
- Select Environment dataset.
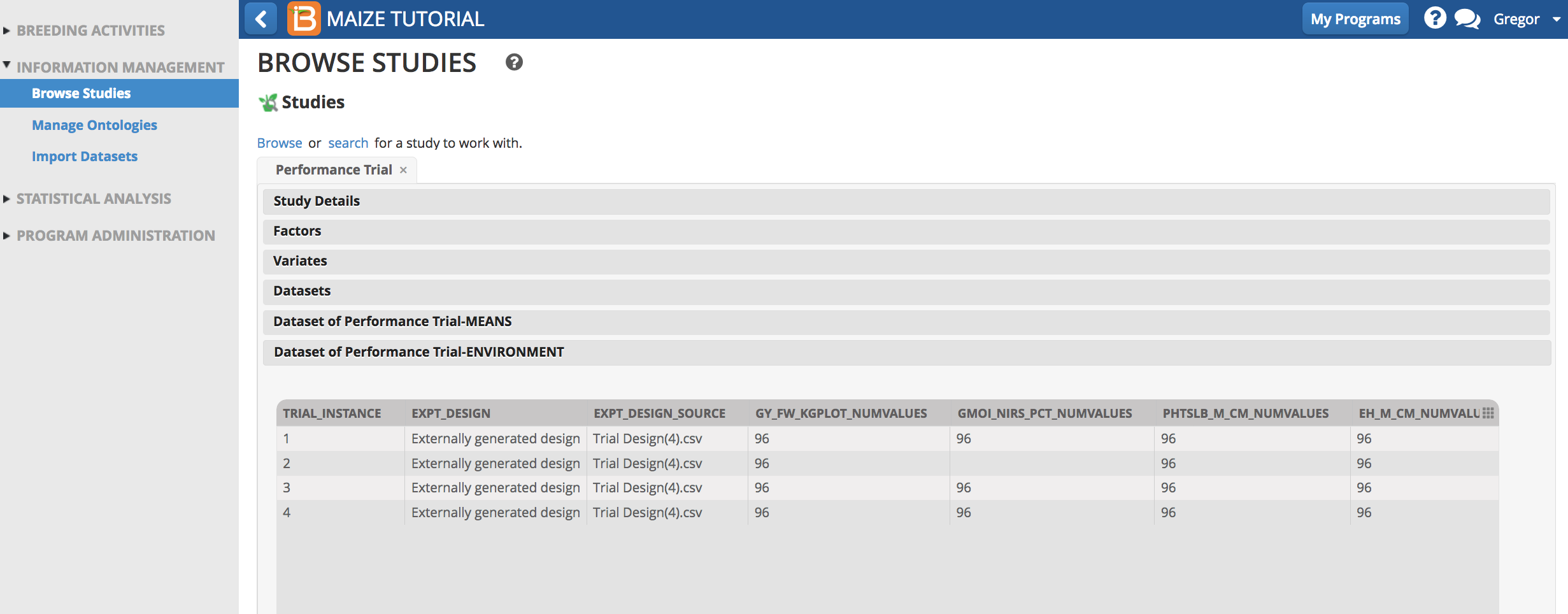
Notice that the environment summary statistics are available for review.
References
Oakey, H., Verbyla, A. P., Pitchford, W., Cullis, B., & Kuchel, H. (2006). Joint modeling of additive and non-additive genetic line effects in single field trials. Theoretical and Applied Genetics, 113, 809–819.
Murray, D. Payne, R, & Zhang, Z. (2014) Breeding View, a Visual Tool for Running Analytical Pipelines: User Guide. VSN International Ltd. (.pdf) (Sample data .zip)
Related Materials
Manual: Manage Trials
Maize: Multi-Site (GxE) Analysis
Funding & Acknowledgements
The Integrated Breeding Platform (IBP) is jointly funded by: the Bill and Melinda Gates Foundation, the European Commission, United Kingdom's Department for International Development, CGIAR, the Swiss Agency for Development and Cooperation, and the CGIAR Fund Council. Coordinated by the Generation Challenge Program the Integrated Breeding Platform represents a diverse group of partners; including CGIAR Centers, national agricultural research institutes, and universities.
The statistical algorithms in the Breeding View were developed by VSN International Ltd in collaboration with the Biometris group at University of Wageningen. Maize demonstration data was provided by Mike Olsen from the CIMMYT, the International Center for Maize and Wheat Improvement, breeding program. These data have been adapted for training purposes. Any misrepresentation of the raw breeding data is the solely the responsibility of the IBP.

This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License