About
This functionality allows users to identify and sample individual plants within a plot for SNP genotyping. . Samples are giving unique IDs that are used to match genotype data to the plant and plot from which it was sampled.
Create Samples within Studies
- From a study where selections have been made, choose 'Create sample list' from the 'Plant level options' of the Actions menu
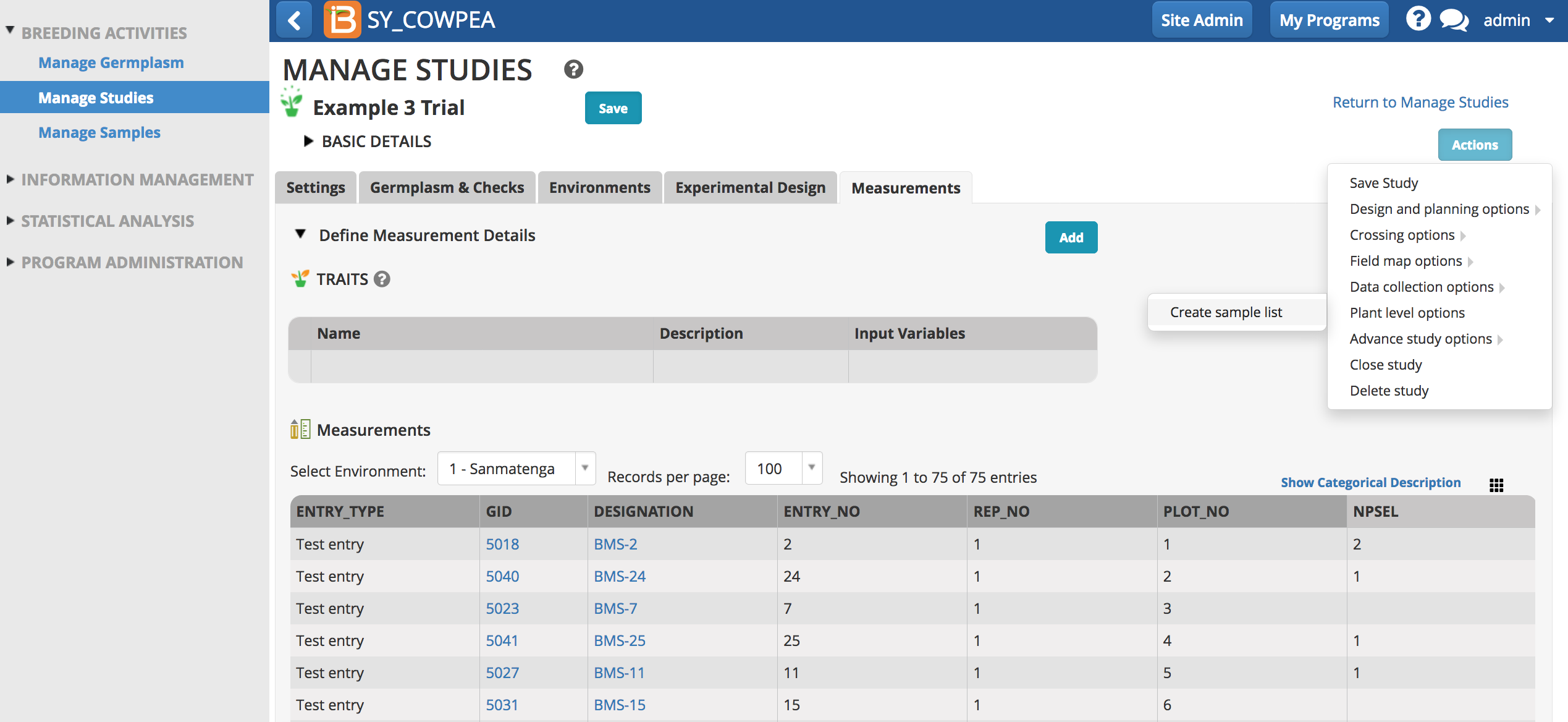
In this example NPSEL (number of plants selected) indicates how many plants from each plot will be sampled.
- Choose the instances you would like to sample.
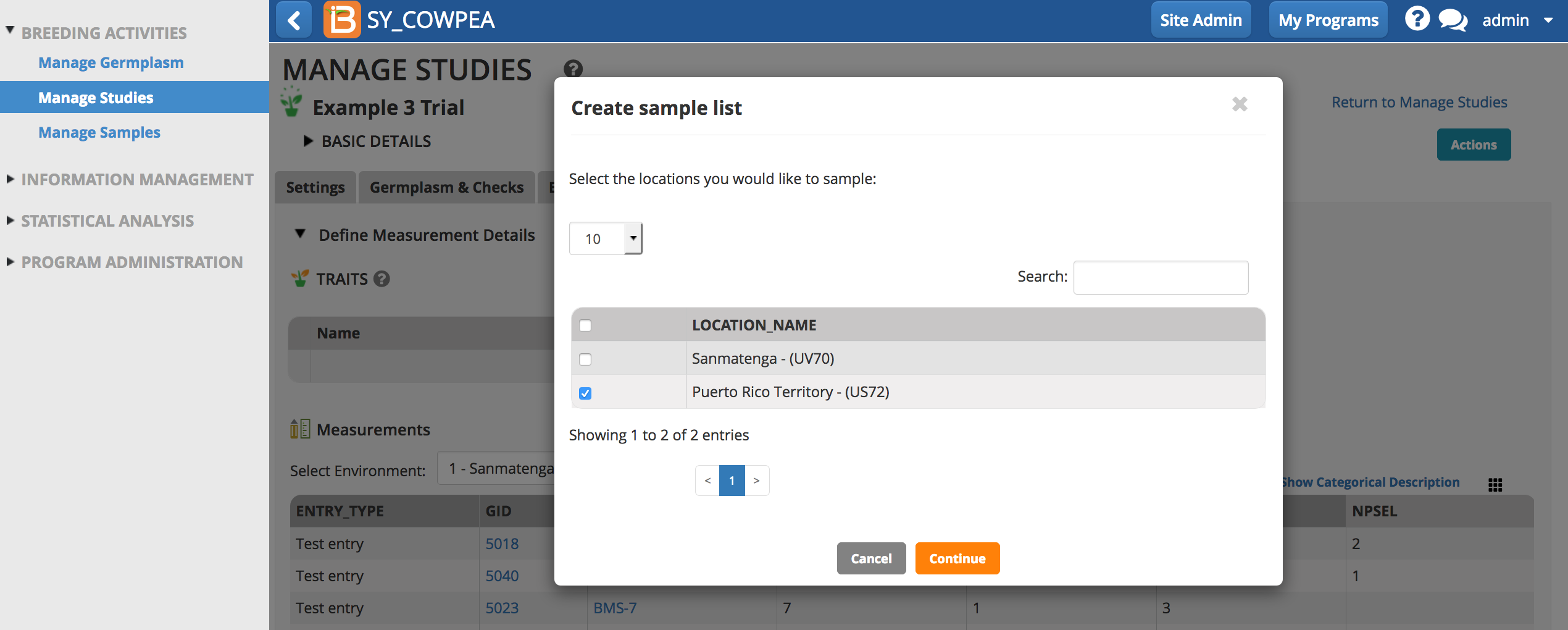
- Choose the variate that defines the number of samples (in this case NPSEL), and enter the optional details about collector and date.
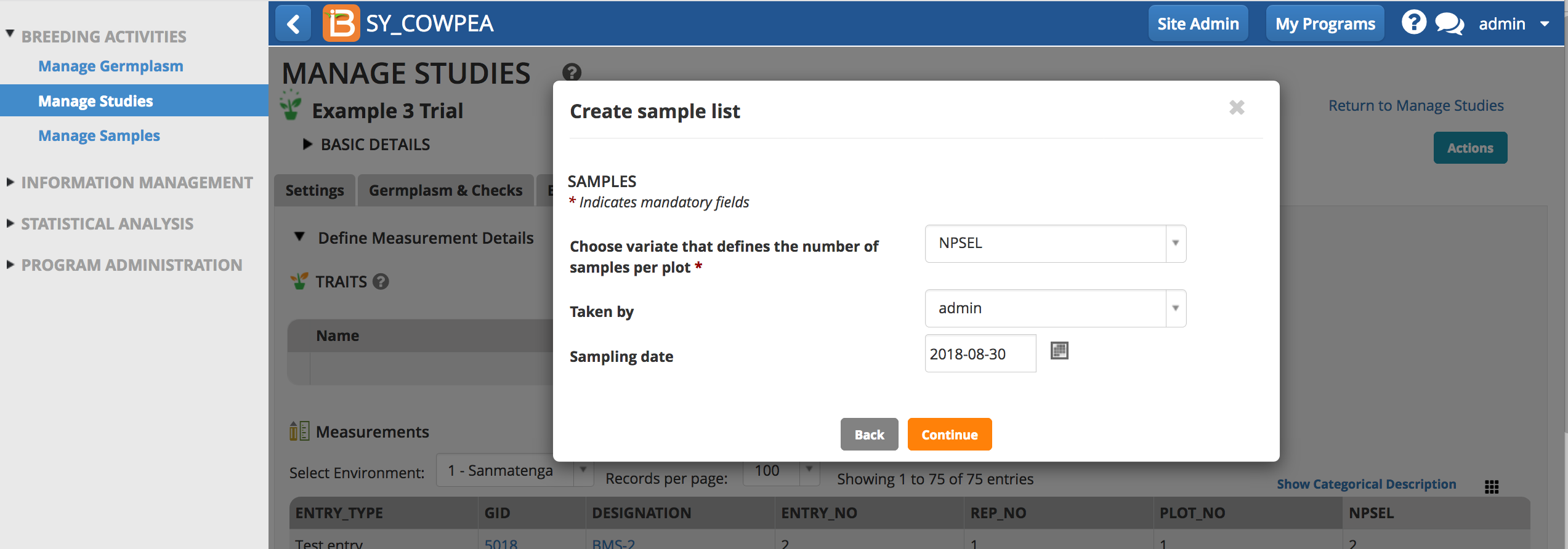
- Save the sample list.
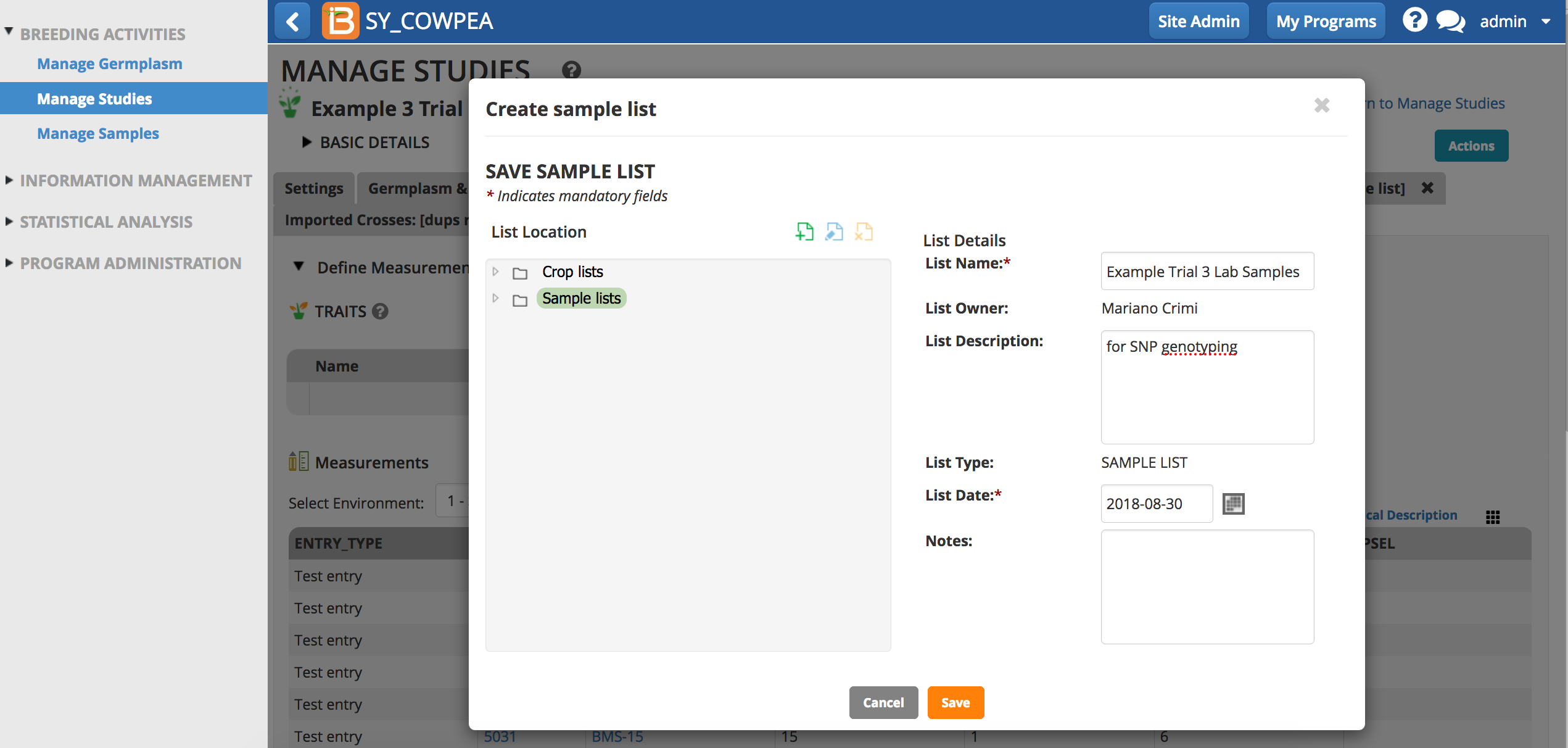
The sample list is now available in the associated study and the sample details are available with other germplasm details.
- Select any germplasm designation link.
Review sample details in germplasm details pop-up.
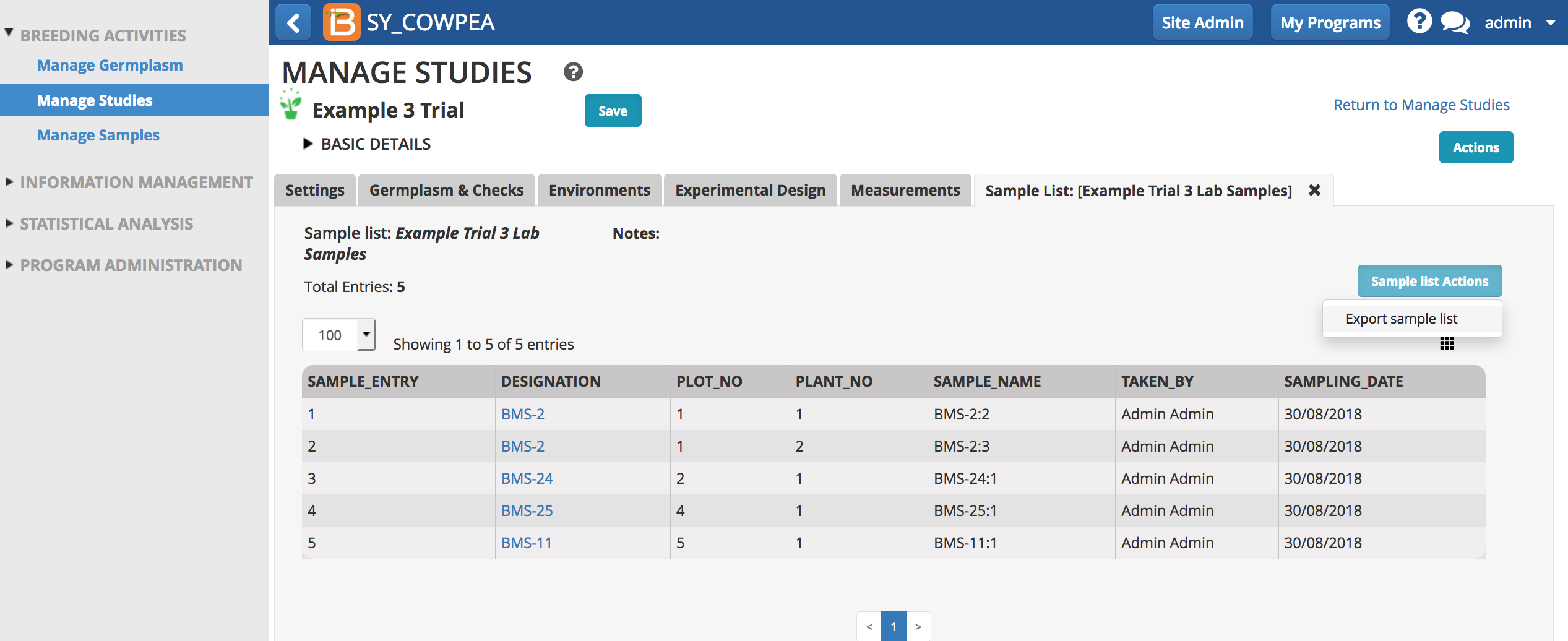
- Export the sample list as an .xls file.

- Review the sample list in Excel. Sample_NAME and SAMPLE_UID will be needed for the eventual upload of genotyping data.
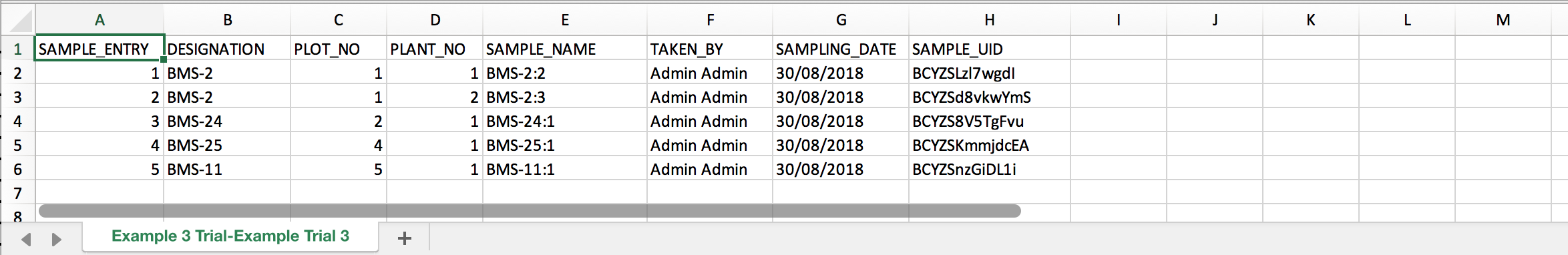
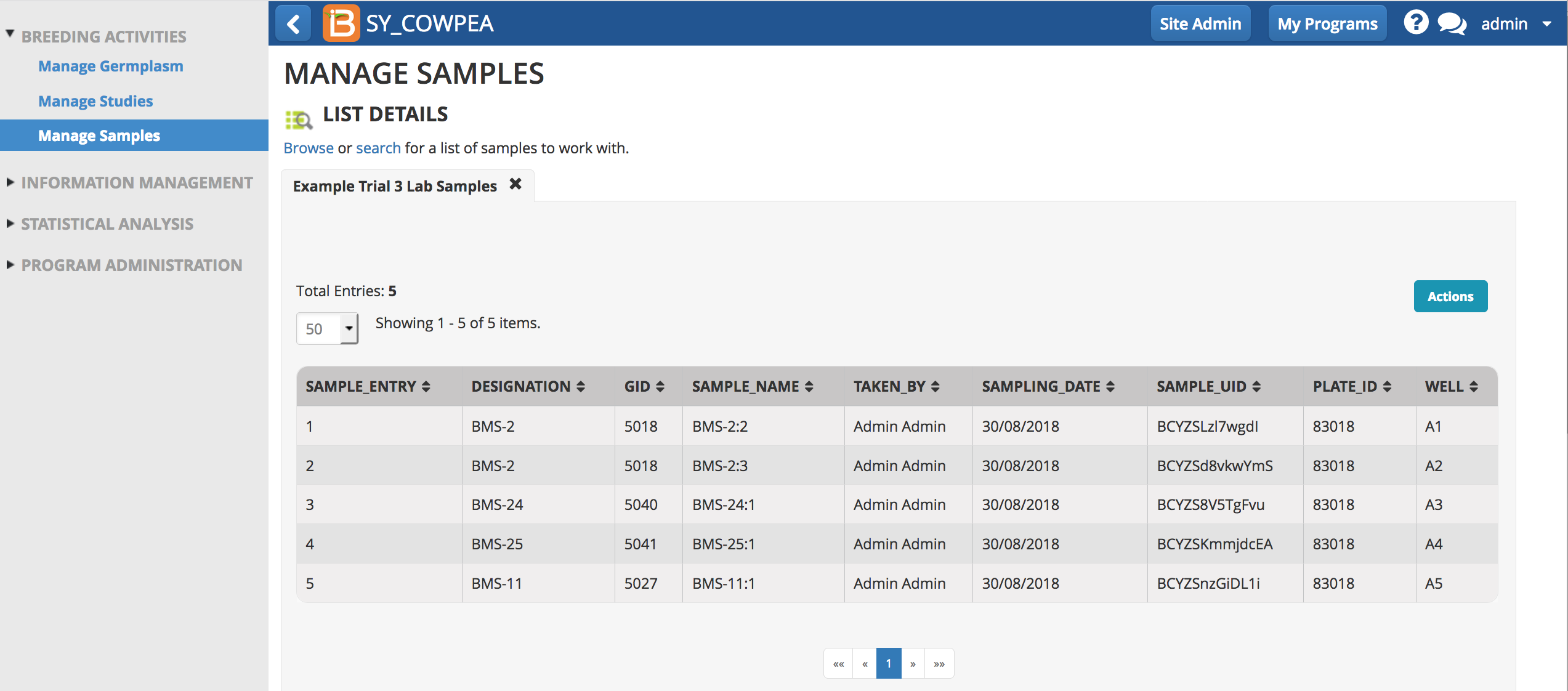
Sample information can be reviewed for individual germplasm from any list in the system. Links from the sample table take the user to the study or dataset.
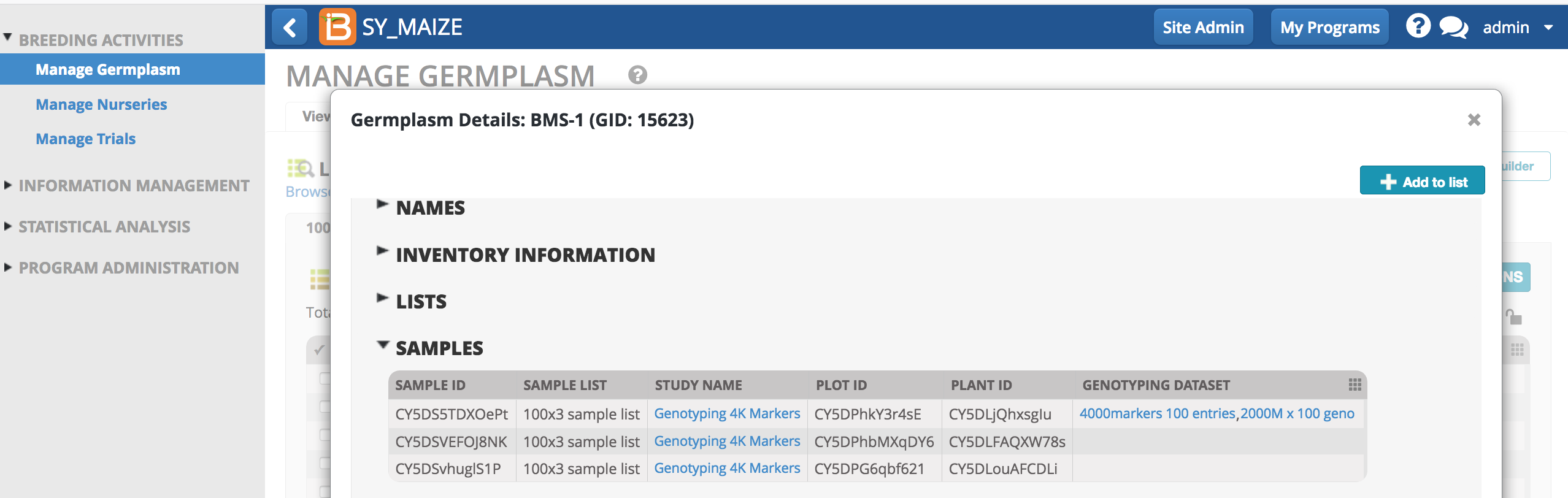
In this example, 3 plants belonging to germplasm BMS-1 have been sampled. The first plant was genotyped twice, once with a set of 4000 markers and once with a set of 2000 markers.
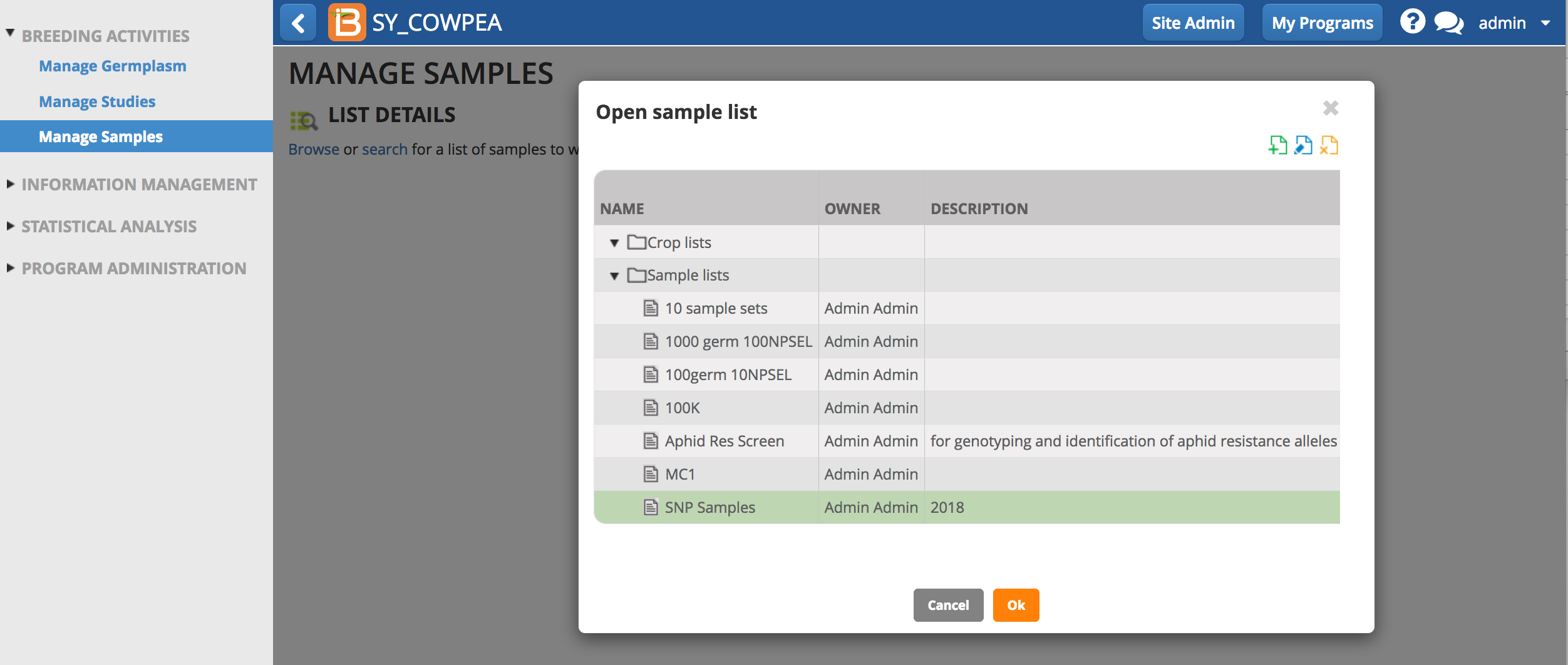
Manage Samples
Browse or search for sample lists.

Browse
- Select Browse. Highlight the list of interest from the directory and select OK.
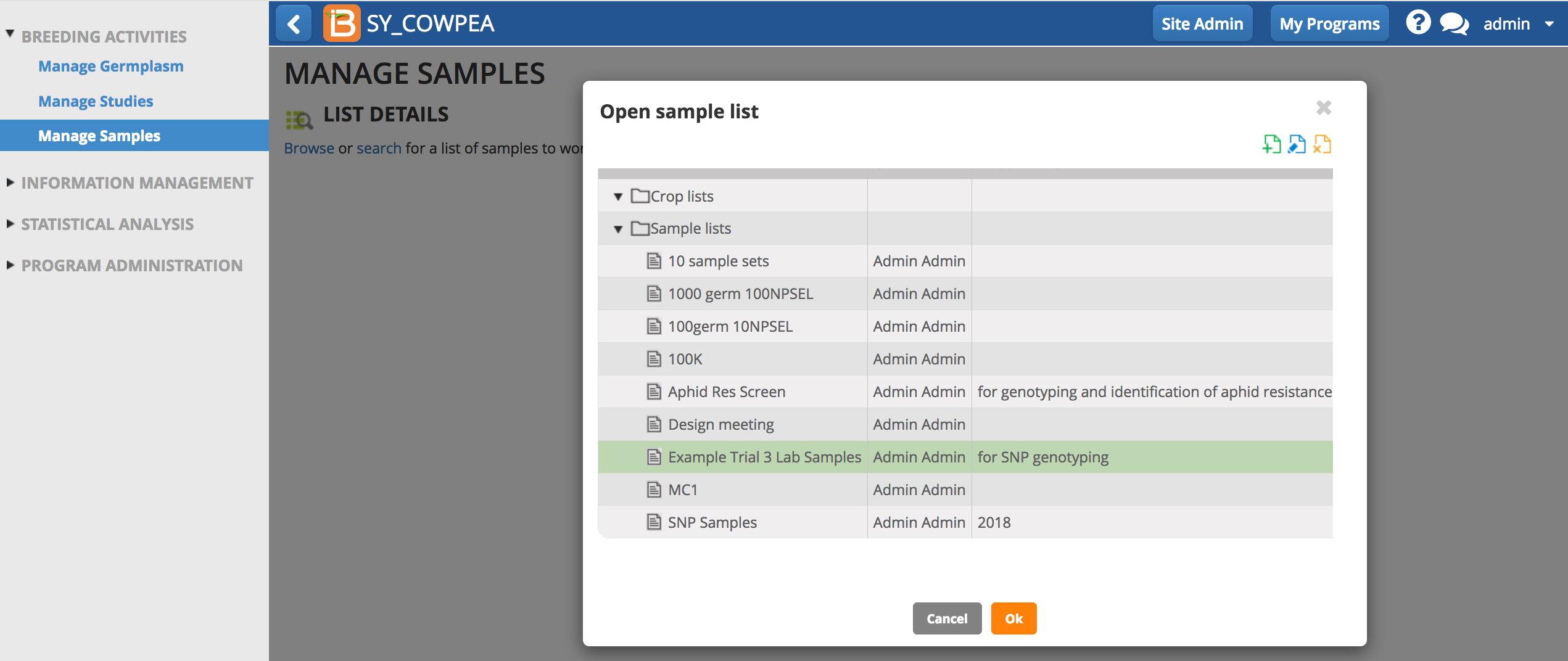
Search
- Enter a partial or exact list name to search the directory. Select the list of interest and close the pop-up.
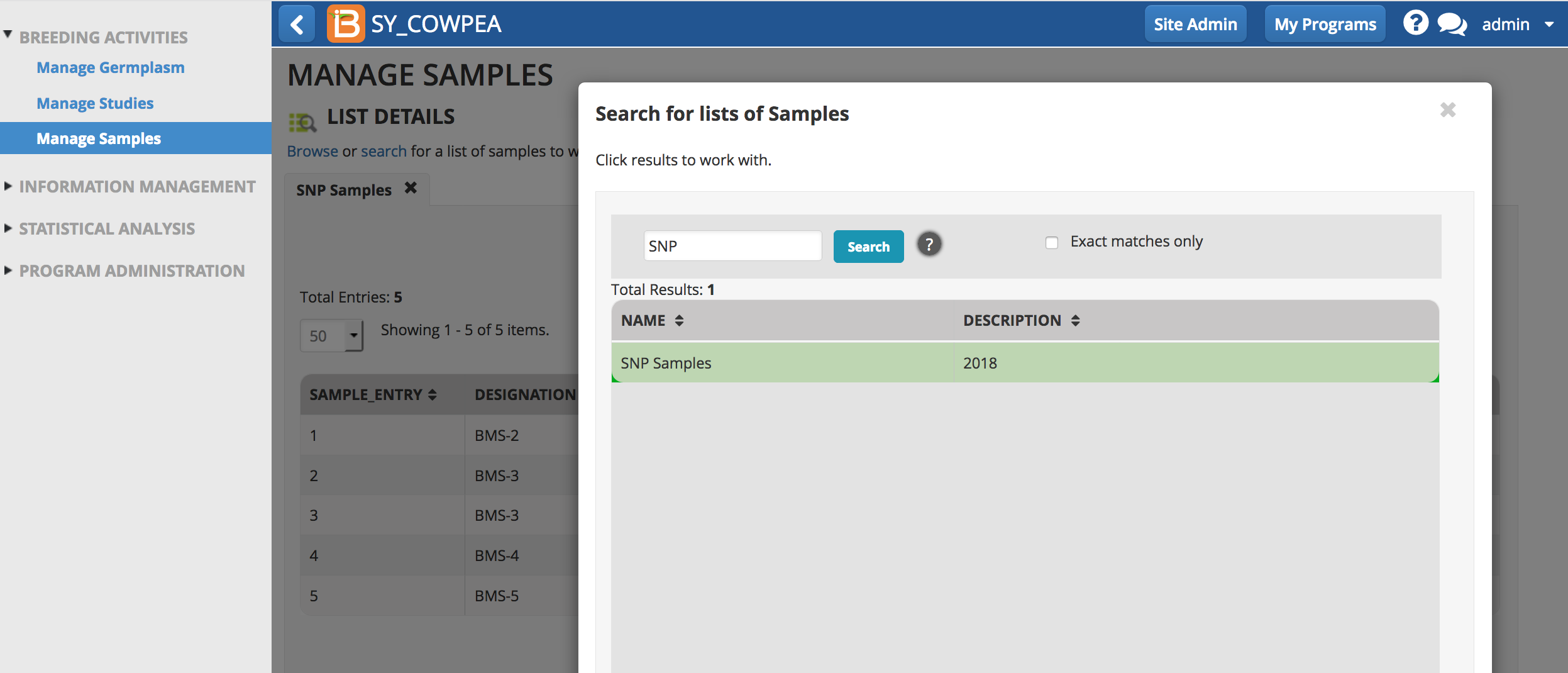
Export Sample List
- Export list (.csv) from the Actions menu.
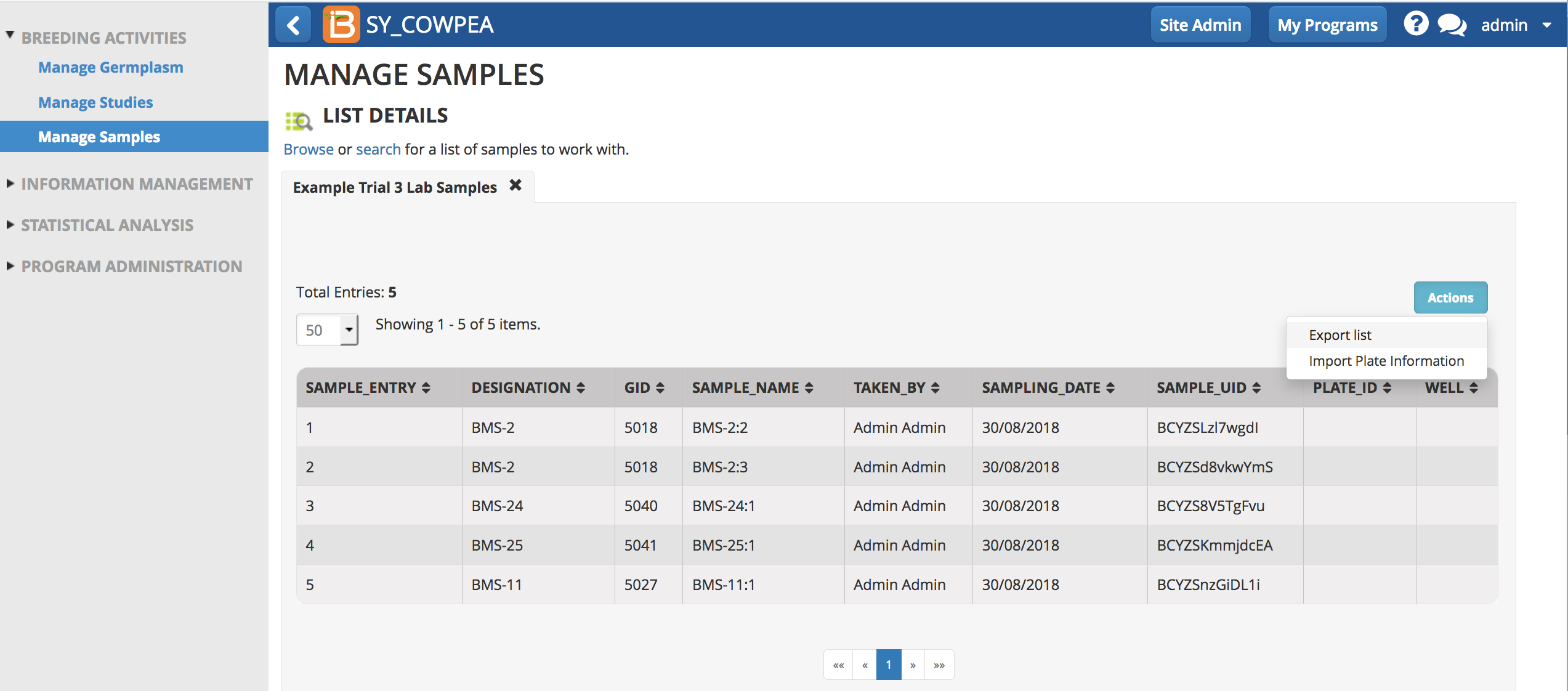
Edit Plate Details
- Open the .csv in a program like Excel. Add columns PLATE_ID and WELL to record plate information. Fill the PLATE_ID and WELL details. Save the .csv file.
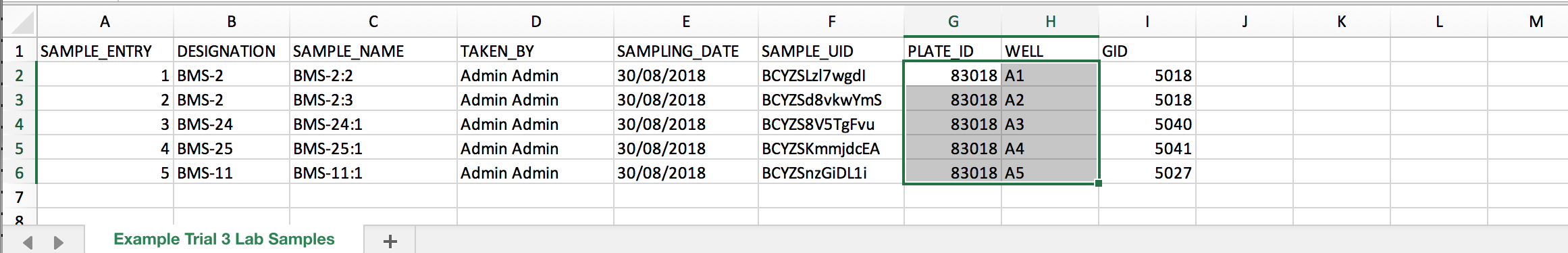
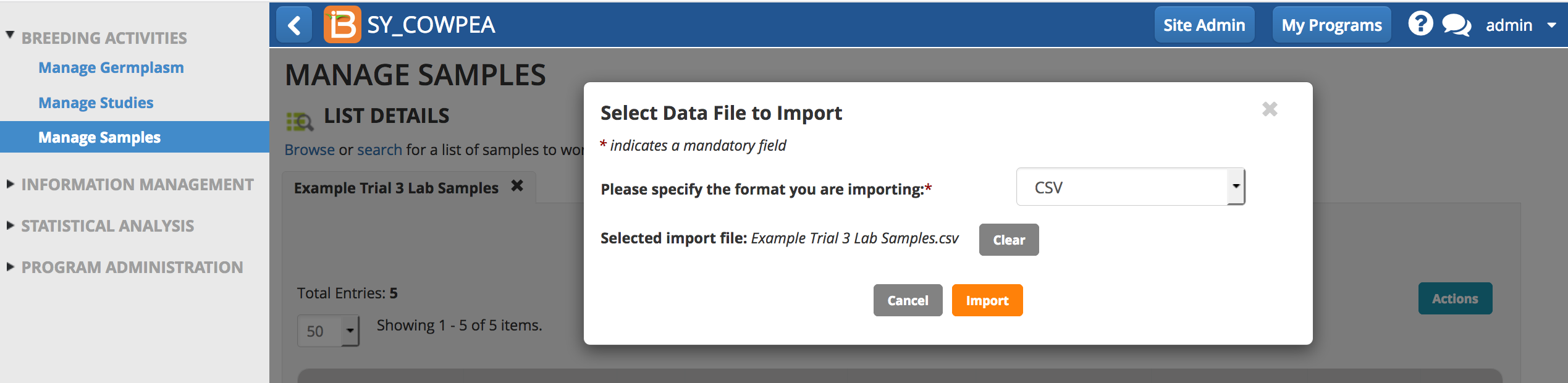
Import Plate Details
- Select .csv file to import.
- Confirm the mapping of column headings and Proceed.
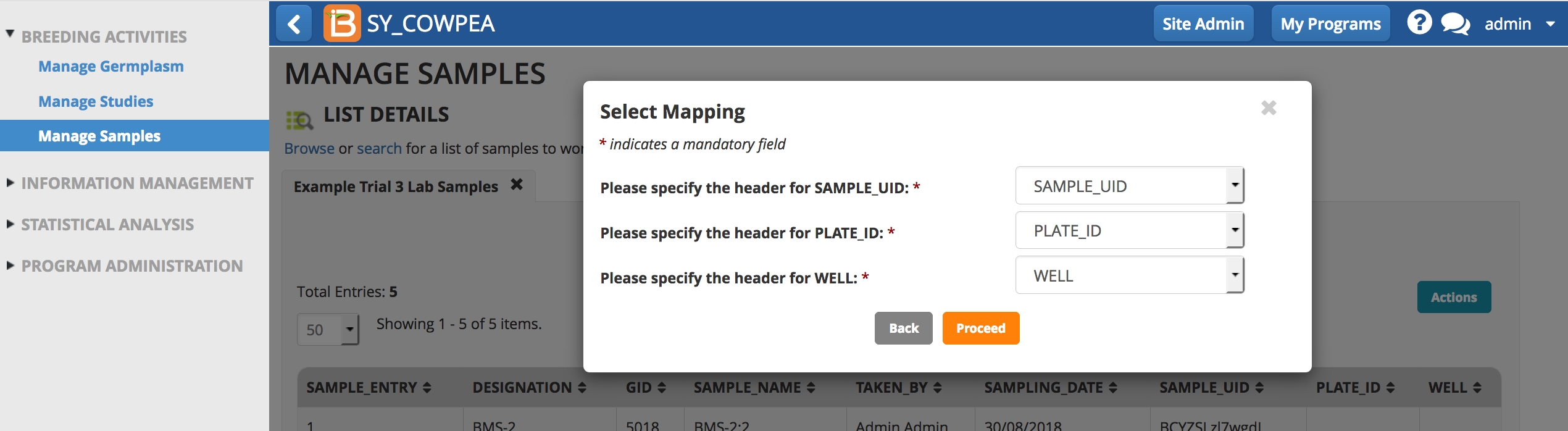
Review the plate details.