About
The dataset importer allows users to upload historical studies from Excel files into the Breeding Management System. There are two ways import datasets, both require completion of preliminary steps to proceed.
Preliminary Steps
Dataset import cannot proceed until preliminary steps are completed.
- Germplasm represented in the dataset must first be imported into to the BMS and a list created.
- All of the phenotypes measured in the dataset must first be defined it the BMS ontology.
- All of the locations represented in the dataset must first be defined in the BMS.
- All of the people represented in the dataset must first be added as BMS users.
Example Data
The following instructions follow import of an example data set after the completion of required preliminary steps.
2014 CIMMYT Maize Study - Start of Import Via Wizard.xls
- 55 maize lines
- Evaluated at 7 locations (instances) across Zimbabwe and Malawi
(1.) CIMMYT Harare
(2.) Art farm Harare
(3.) Glendale
(4.) Chirdezi
(5.) Matopos
(6.) Muzarabani
(7.) Dedza - 2 reps per location
- Randomized complete block design
- 20 phenotypes measured
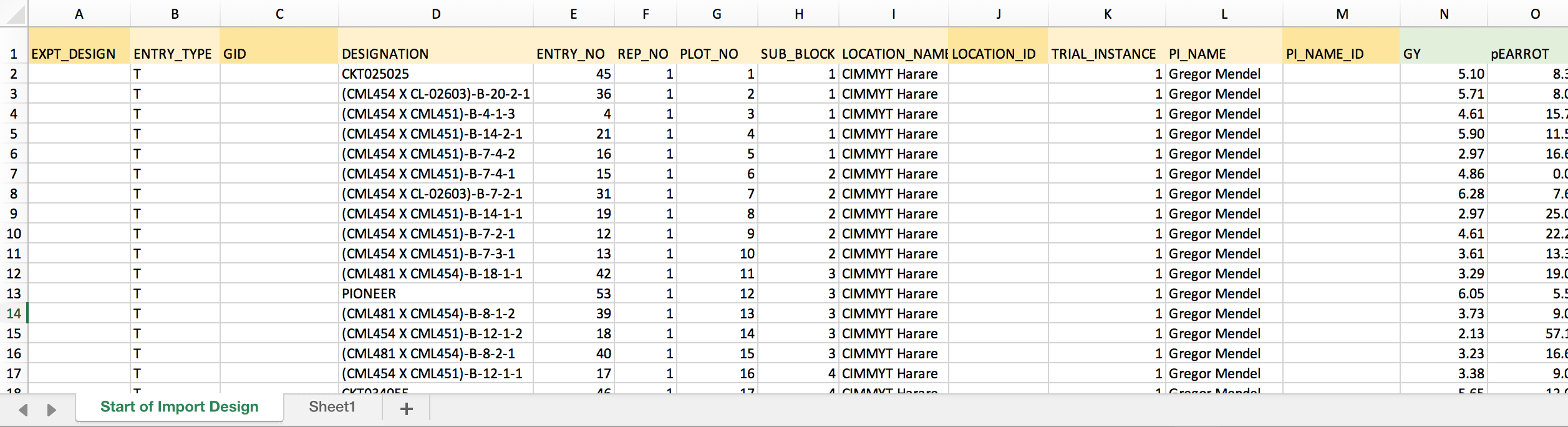
The example dataset: Gold colored columns of data represent those that should be formatted before import. The darker gold columns are BMS-specific indentifiers that must be matched before import.
Data Import Wizard
The Data Import Wizard is the most flexible way to import historical data from trials and nurseries. Some critical identifiers must be perfectly formatted, but other data, like phenotypic names can be mapped to BMS-specific terms. Once a mapping is completed, the BMS "remembers" past mapping to ease subsequent dataset uploads.
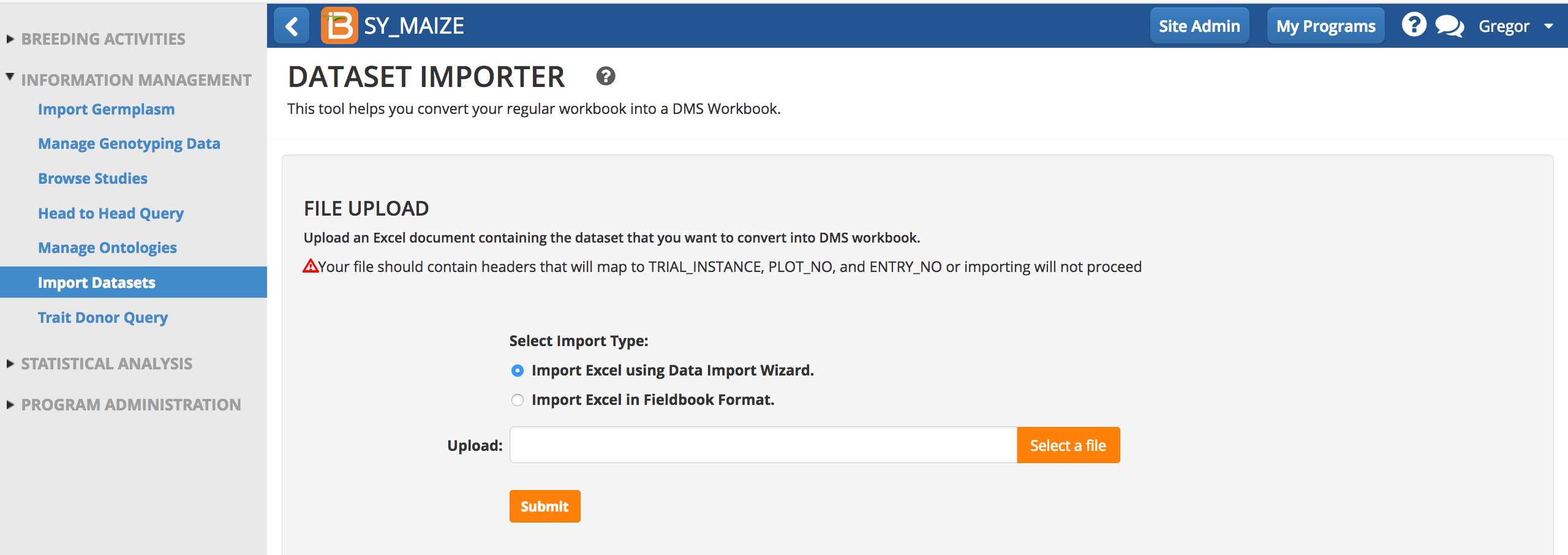
Excel File Format
The Wizard will considers one worksheet of observation data (.xls). Formatting and some data matching is needed for trial design and BMS identifiers. The phenotypes do not require any formatting. Three columns of data are required to import phenotypic observations: TRIAL_INSTANCE, ENTRY_NO, and GID. These column headers must be exact. However you will probably also want to include additional descriptive data.
| Common Columns of Data | About |
|---|---|
| EXPT_DESIGN | Needed to be included and filled for the study to be available from Manage Studies
|
| ENTRY_TYPE | Needed to differentiate test entries from checks
|
| GID | REQUIRED to match germplasm to the BMS database. Can be found in BMS Manage Germplasm. Open the germplasm list, lock it, and export as .xls file. Importing this list into the BMS is one the required preliminary steps (see above). |
| DESIGNATION | Human readable germplasm name. |
| ENTRY_NO | REQUIRED sequential numbering of the germplasm list (1,2,3.....,n). |
| REP_NO | Sequential numbering of replicates per instance (1,2,3.....,n). |
| PLOT_NO | Sequential numbering of plots per instance (1,2,3.....,n). |
| SUB_BLOCK | Sequential numbering of blocks per instance (1,2,3.....,n). |
| LOCATION_NAME | Human readable location name. |
| LOCATION_ID | Needed to match locations to the BMS database. Can be exported directly from the database tables or read from the Manage Program Settings>Locations interface, |
| TRIAL_INSTANCE | REQUIRED sequential numbering of the instances per study (1,2,3.....,n). Generally there is a one-to-one relationship between instance and location, but instance can also represent time. For example, a single location can be associated with two instances of study that took place over two \years. |
| PI_NAME | First and last name of a BMS user |
| PI_NAME_ID | Needed to match users to the BMS database. Can be exported directly from the database tables or read from the description sheet (.xls) of study created by the person of interest. |
Simple Data Matching
EXPT_DESIGN
- The example dataset has a randomized complete block design. Enter "RCBD" into the first empty cell of the column. Copy.
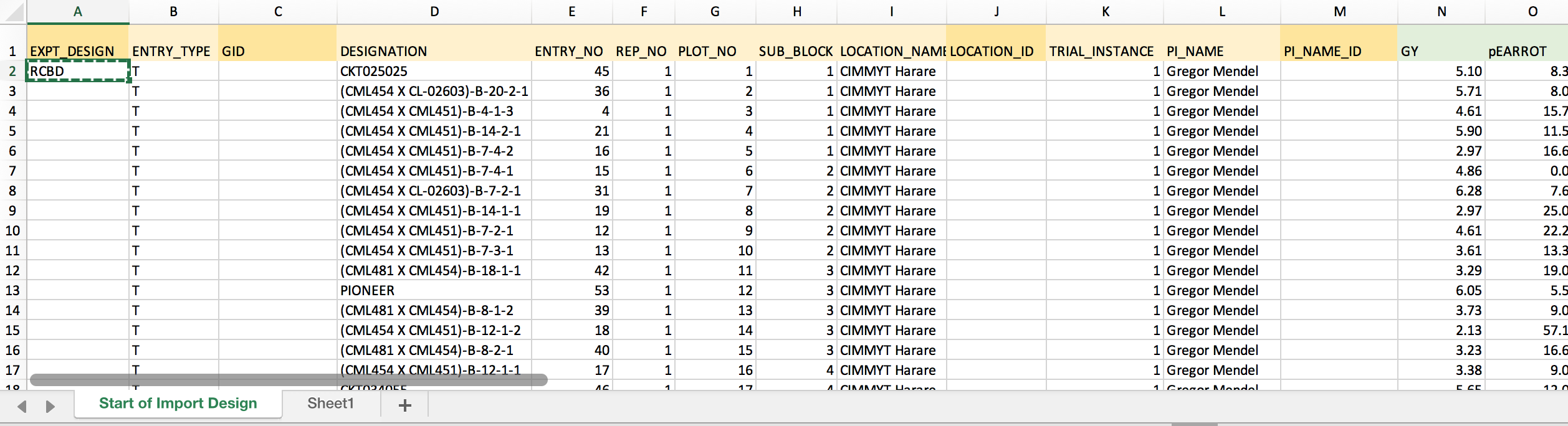
- Paste to fill all 770 cells of the column with "RCBD".
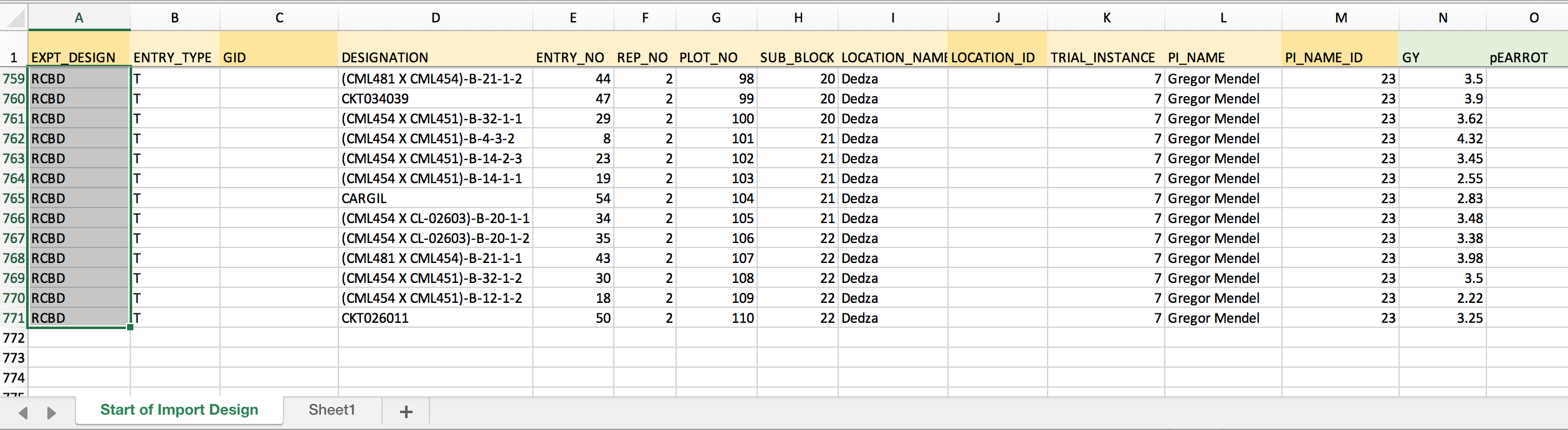
PI_NAME_ID
- There is only one person associated with this dataset, Gregor Mendel. His PI_NAME_ID in the example BMS is 23. Enter PI_NAME_ID in the first empty column. Copy. Paste to fill all 770 cells of the column with PI_NAME_ID.
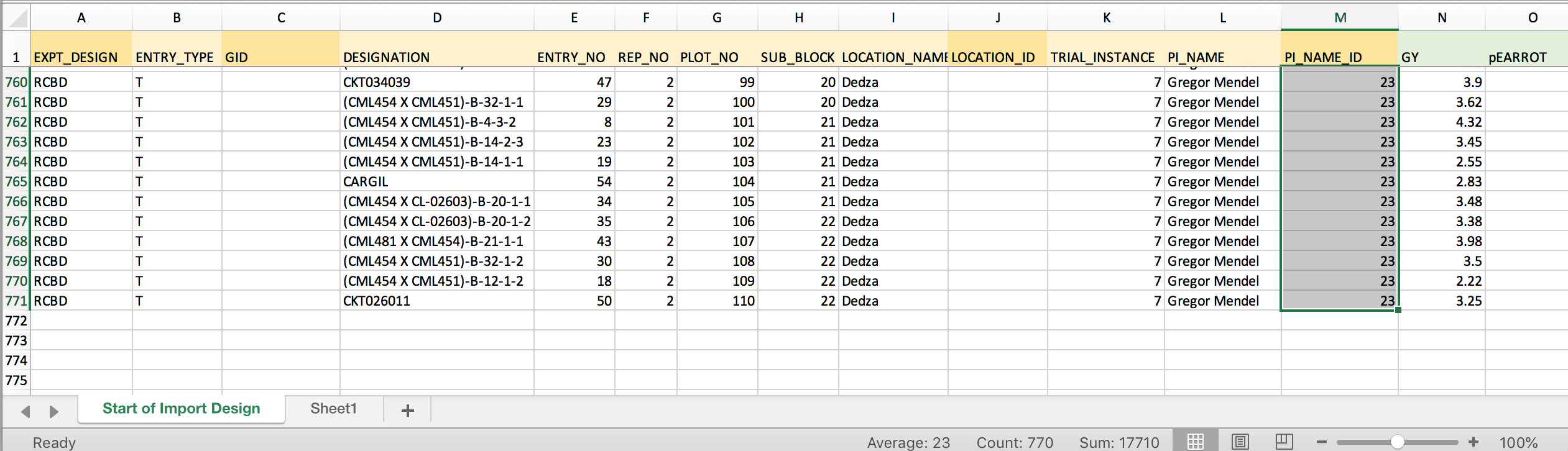
Matching with VLOOKUP
In columns where there isn't a one-to-one relationship for data matching you can use a matching array and VLOOKUP to match data.
- Open a second worksheet. Create matching arrays for the 55 germplasm and their GIDs as well as the 7 TRIAL_INSTANCEs and their Location_IDs.
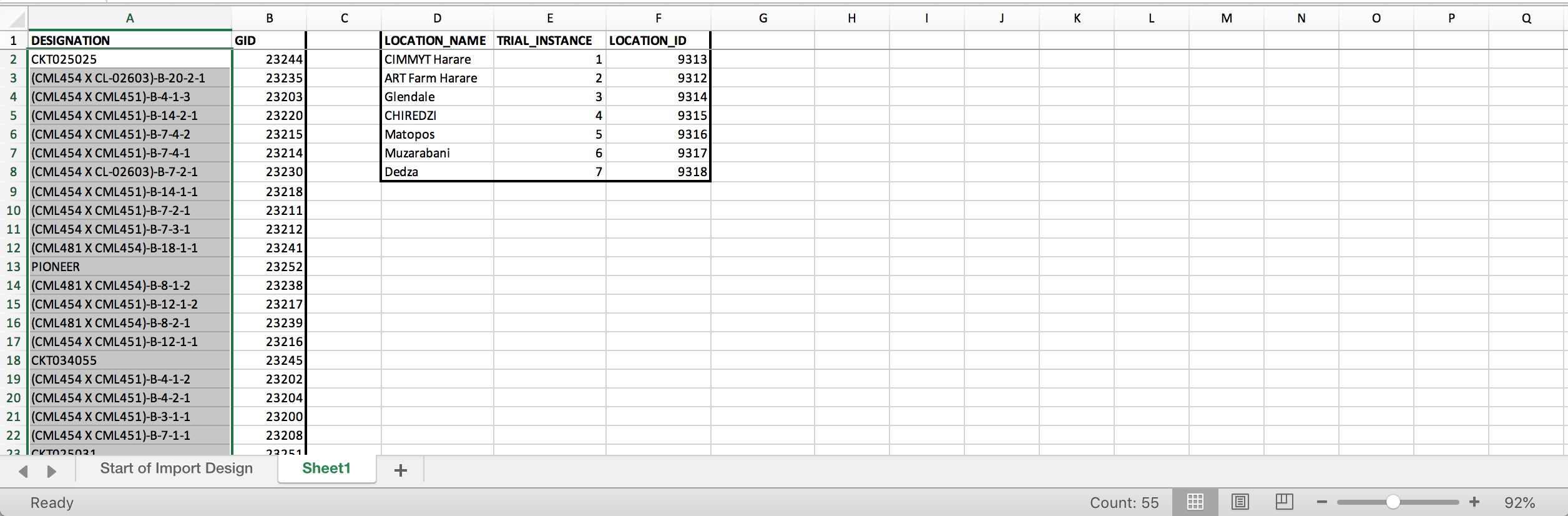
Two data matching arrays. The germplasm array contains columns of data copied from a BMS list export file. The location array was created manually by matching location, instance, and LOCATION_IDs. LOCATION_IDs were copied from the Manage Program Settings-Locations interface.
GID
- Return to the dataset worksheet. Develop the VLOOKUP formula needed to retrieve GID based on DESIGNATION. Lock the coordinates for the matching array by inserting $ before each position.
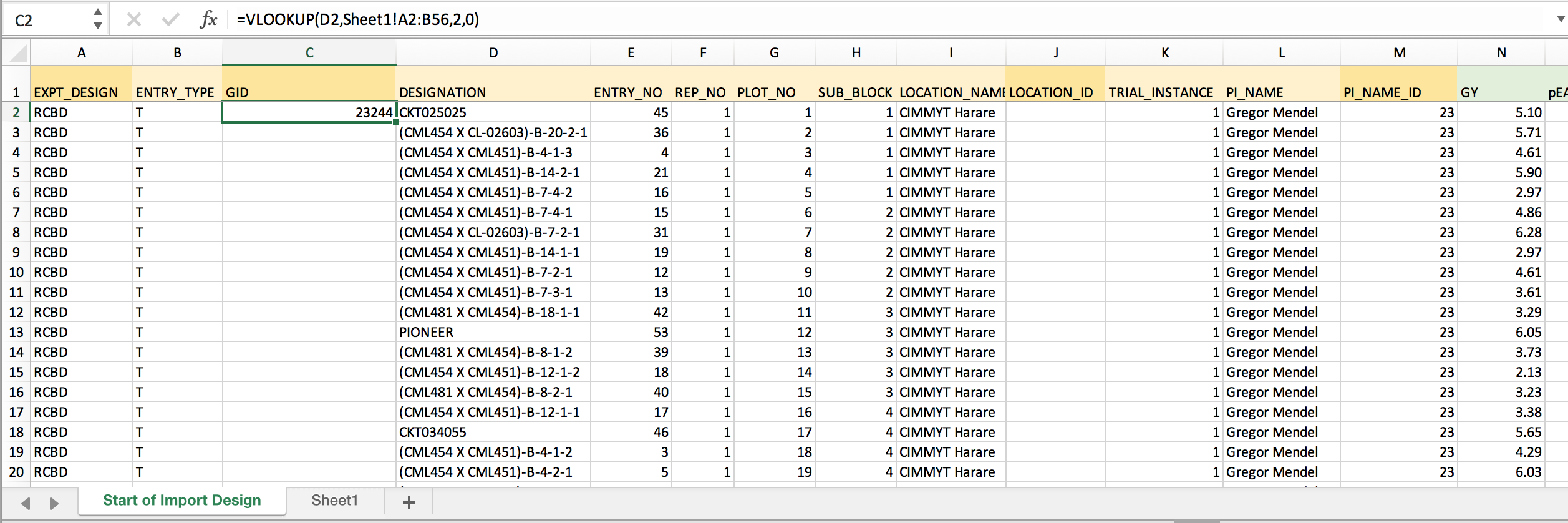
=VLOOKUP(D2,Sheet1!$A$2:$B$56,2,0) Retrieves GID from 2nd column of the matching array based on exact DESIGNATION matches. The dollar signs lock the match array so that the formula can be copied to down the column.
- Copy the formula and paste down the entire column to match all designations with GID.
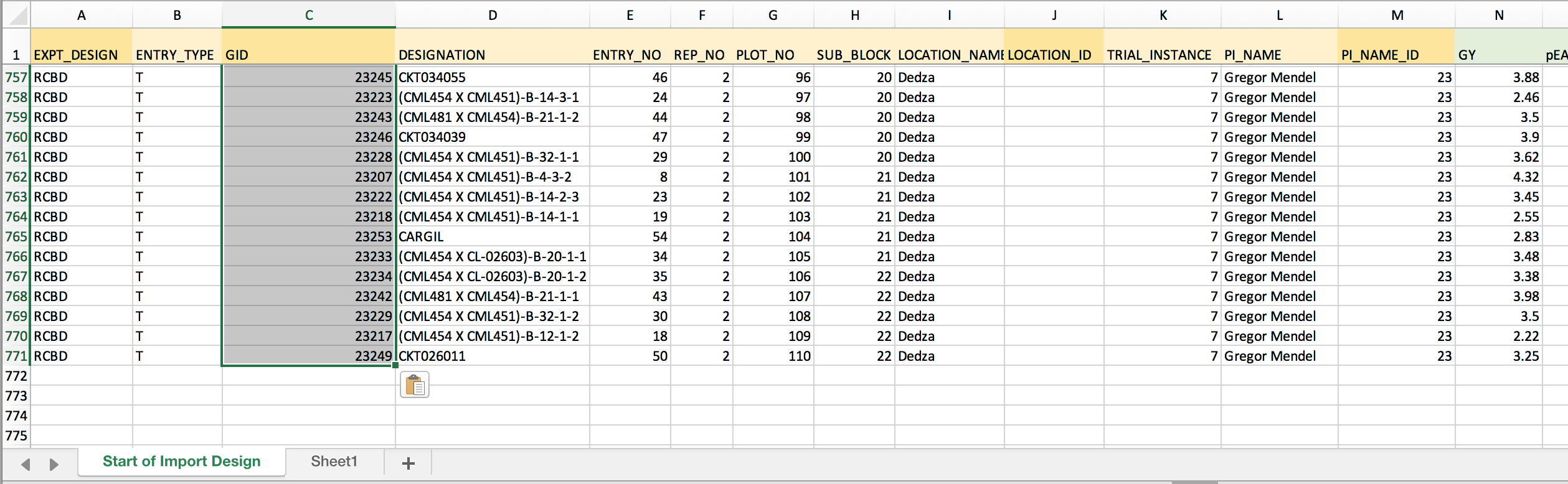
LOCATION_ID
- Develop the VLOOKUP formula needed to retrieve LOCATION_ID based on TRIAL_INSTANCE. Lock the coordinates for the matching array by inserting $ before each position.
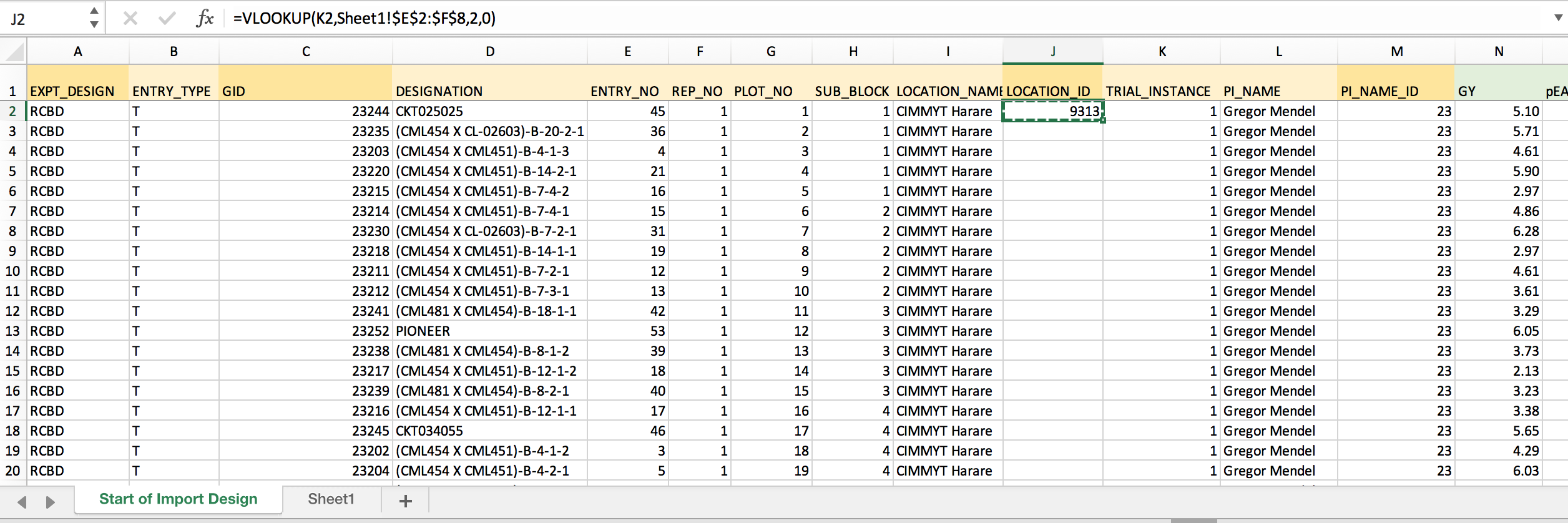
=VLOOKUP(K2,Sheet1!$E$2:$F$8,2,0) Retrieves Location ID from 2nd column of the matching array based on exact TRIAL_INSTANCE matches. The dollar signs lock the match array so that the formula can be copied to down the column.
- Copy the formula and paste down the entire column to match all TRIAL_INSTANCES with LOCATION_ID.
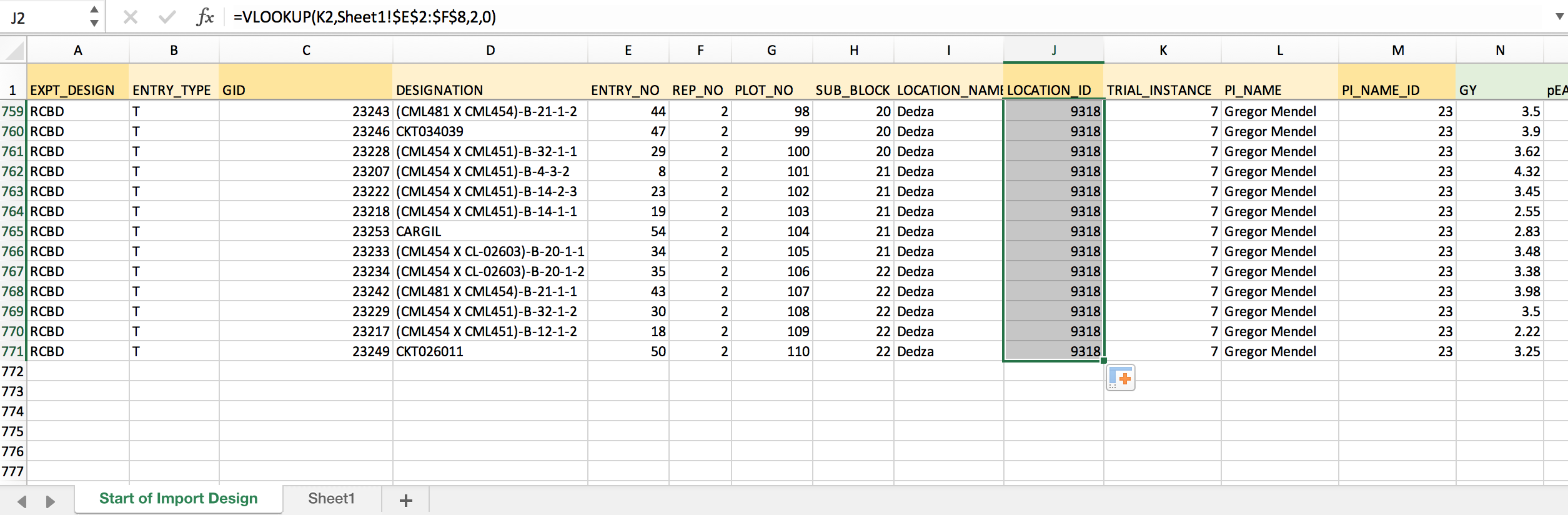
Remove Formulas & Save
The BMS will reject an .xls file with cells that contain formulas.
- You must create empty columns for each colum of data that contains formulas. Copy and paste "As Values" all columns of data that contain formulas. Delete the columns that contain formulas.
- Save the (.xls) file. If you are using a newer version of Excel, you may be asked to confirm the file type.
Start Import
Example File: Ready to Import Dataset (.xls)
- Select the Data Import Wizard, and choose an .xls file to import. Submit.
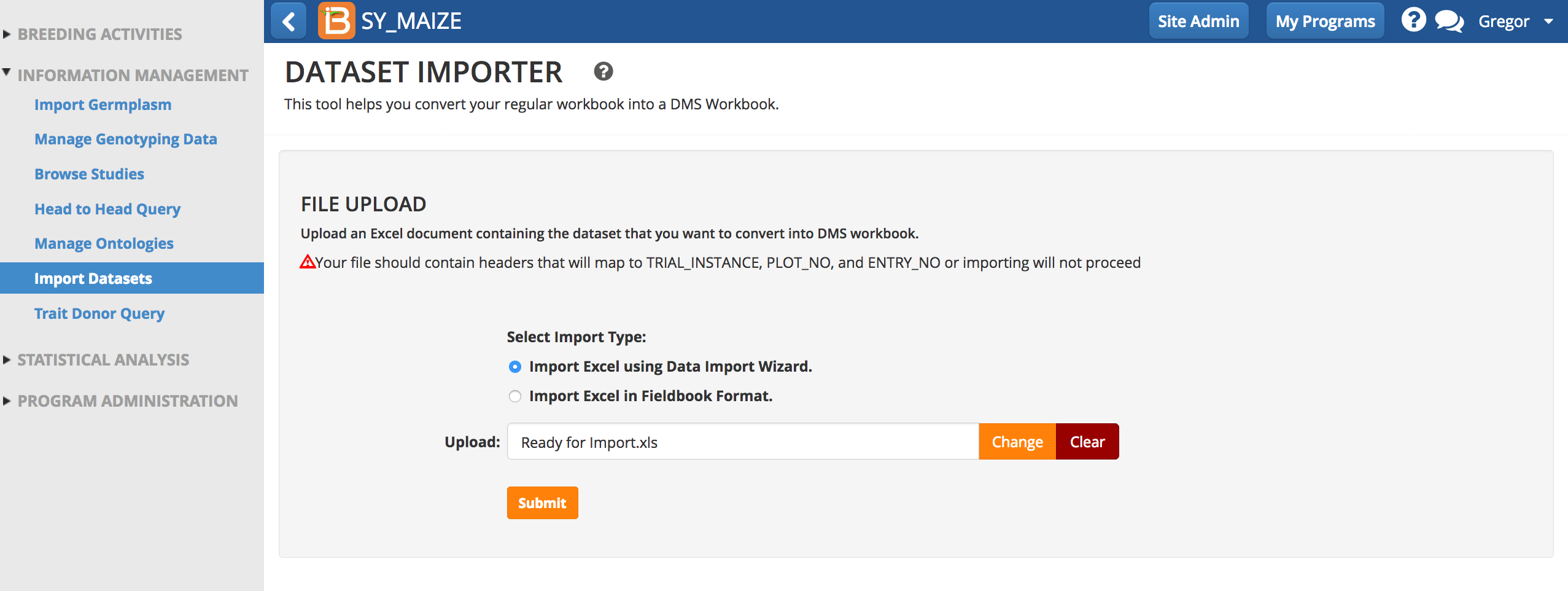
- Specify all of the study details and select Next.
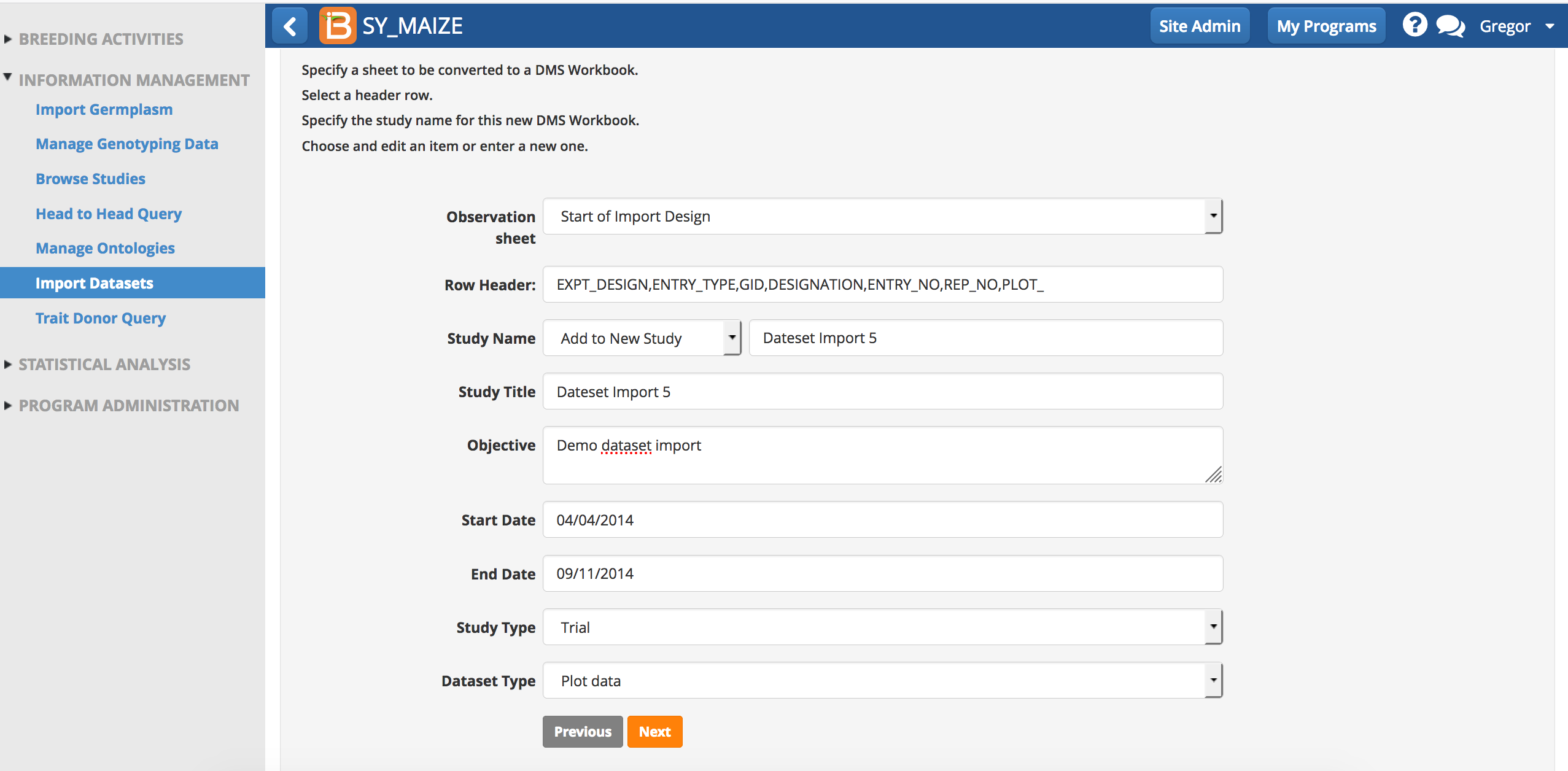
Review & Save Mapping
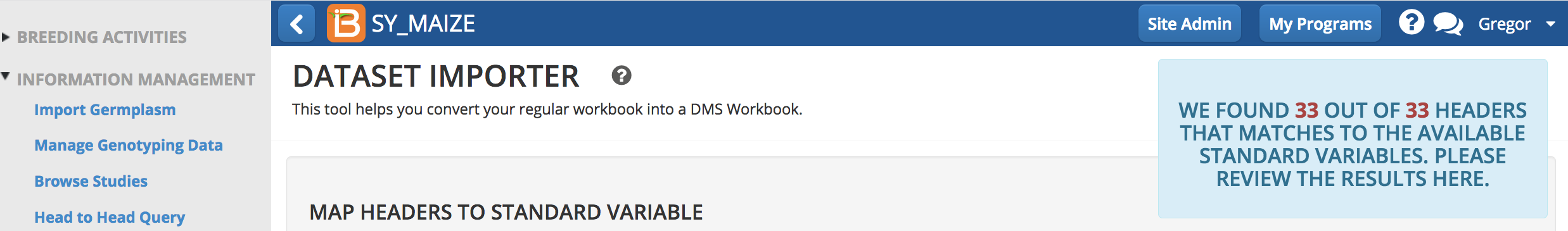
The BMS will attempt to map all the headers in the dataset file. Once a dataset has been mapped, the BMS will rember the mapping.
- Carefully review BMS mapping. If a term is unmatched drag it to the appropriate variable category and match it to the BMS ontology. If a term is mapped incorrectly, use the re-mapping function to find the correct ontology term. If a match term can't be found, data import will need to stop until the term is added to the BMS ontology. Save mapping.
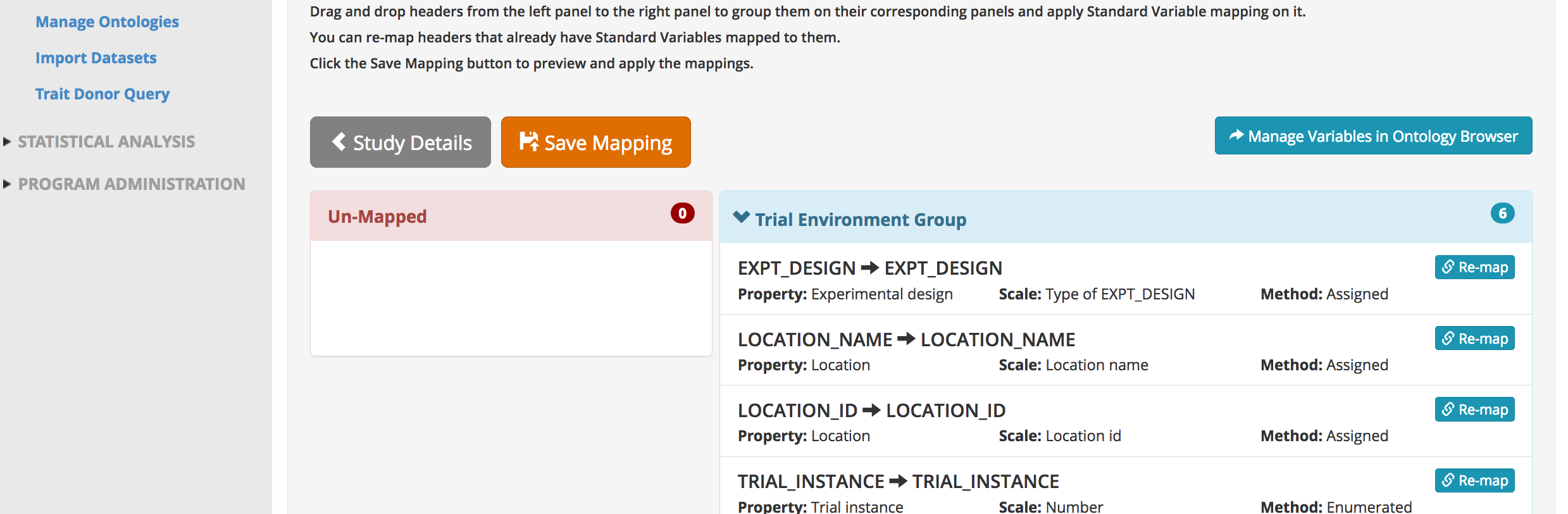
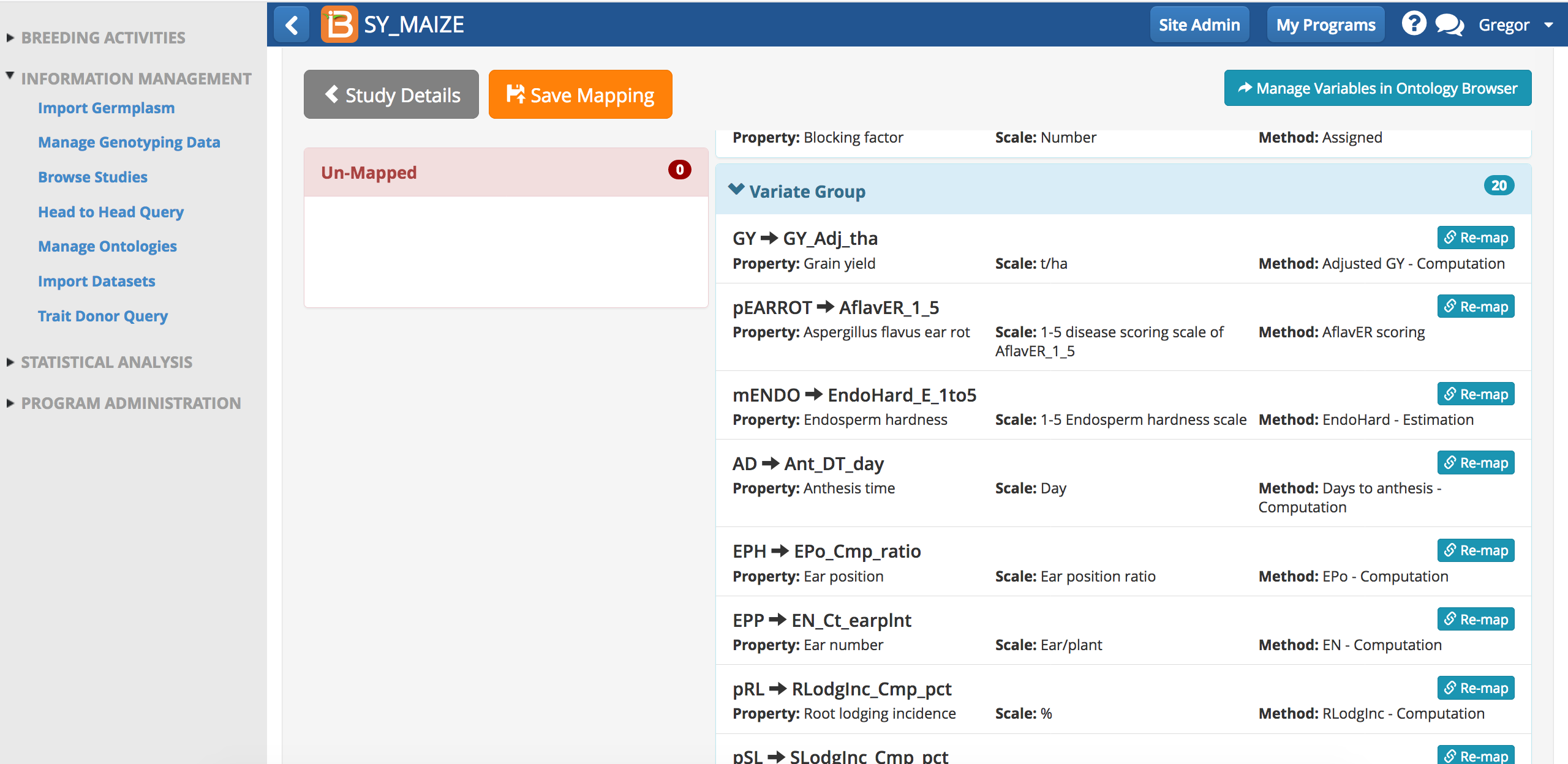
Notice that the names for phenotypes in the dataset differ from the names in the BMS ontology. Since these traits were manually matched previously, the BMS is able to subsequently match this terms without user input.
Correct Mapping Errors
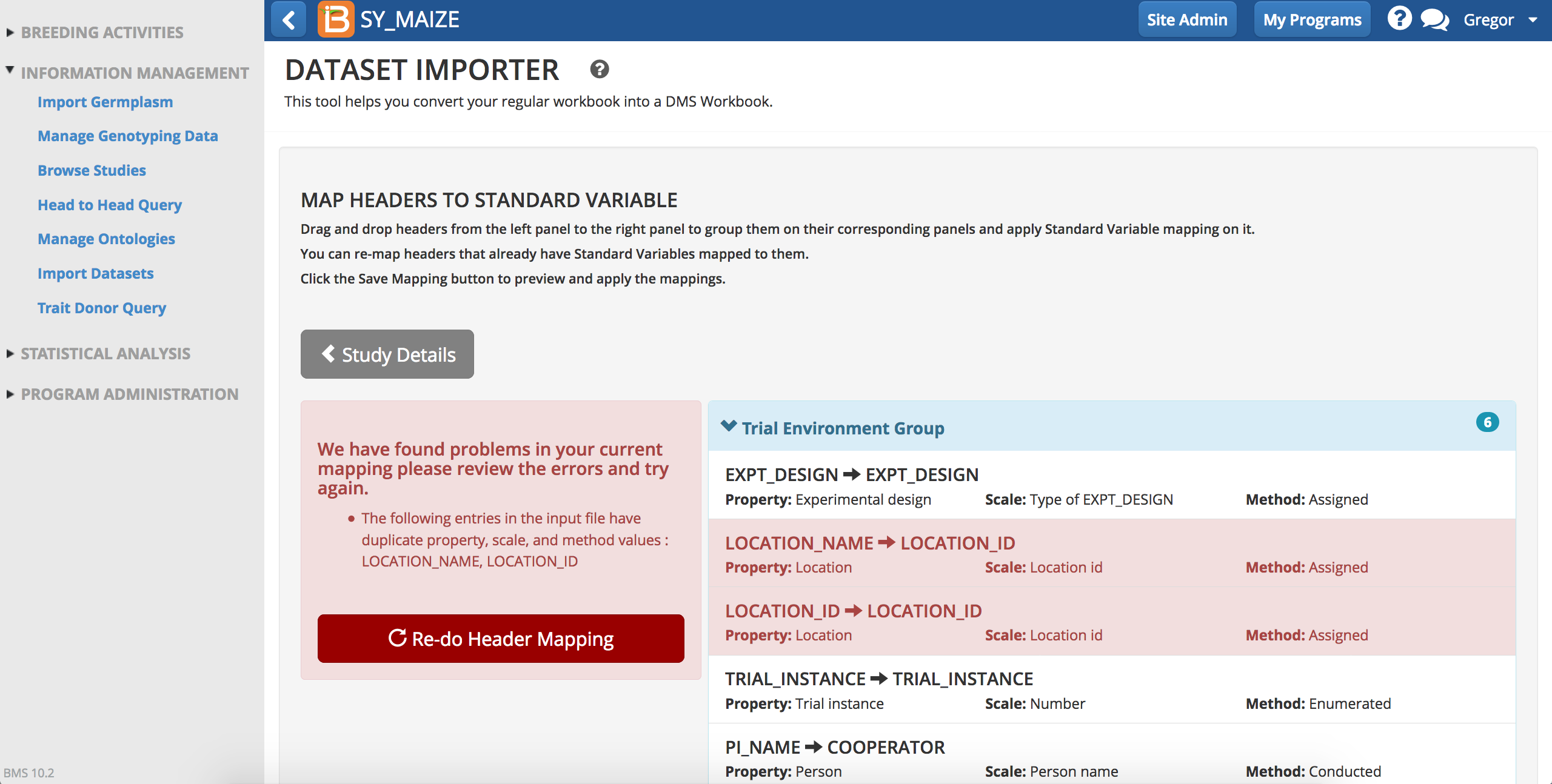
Upon saving, the BMS validates mapping and will flag errors.
- Select Re-do header mapping.
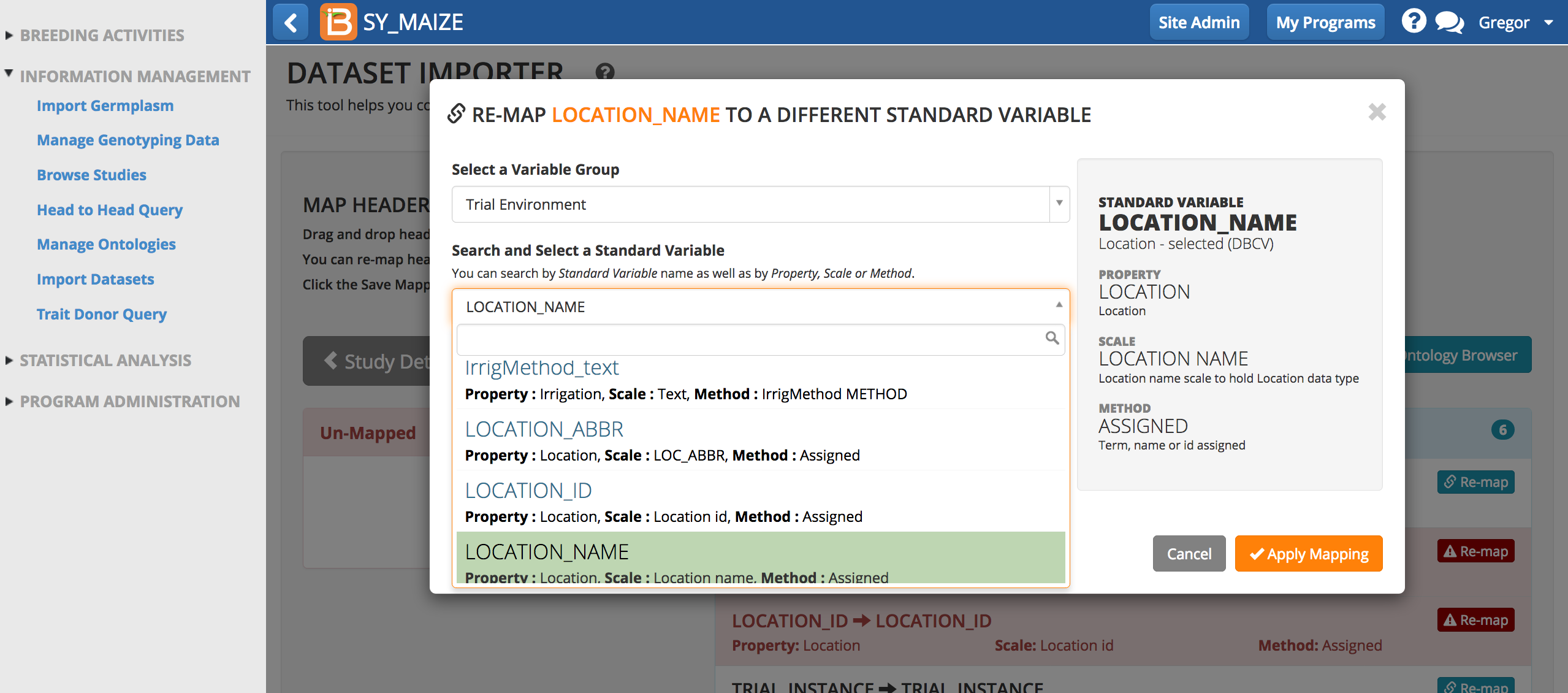
- Remap to the correct term and apply mapping. Save mapping again.
- Confirm header mapping.
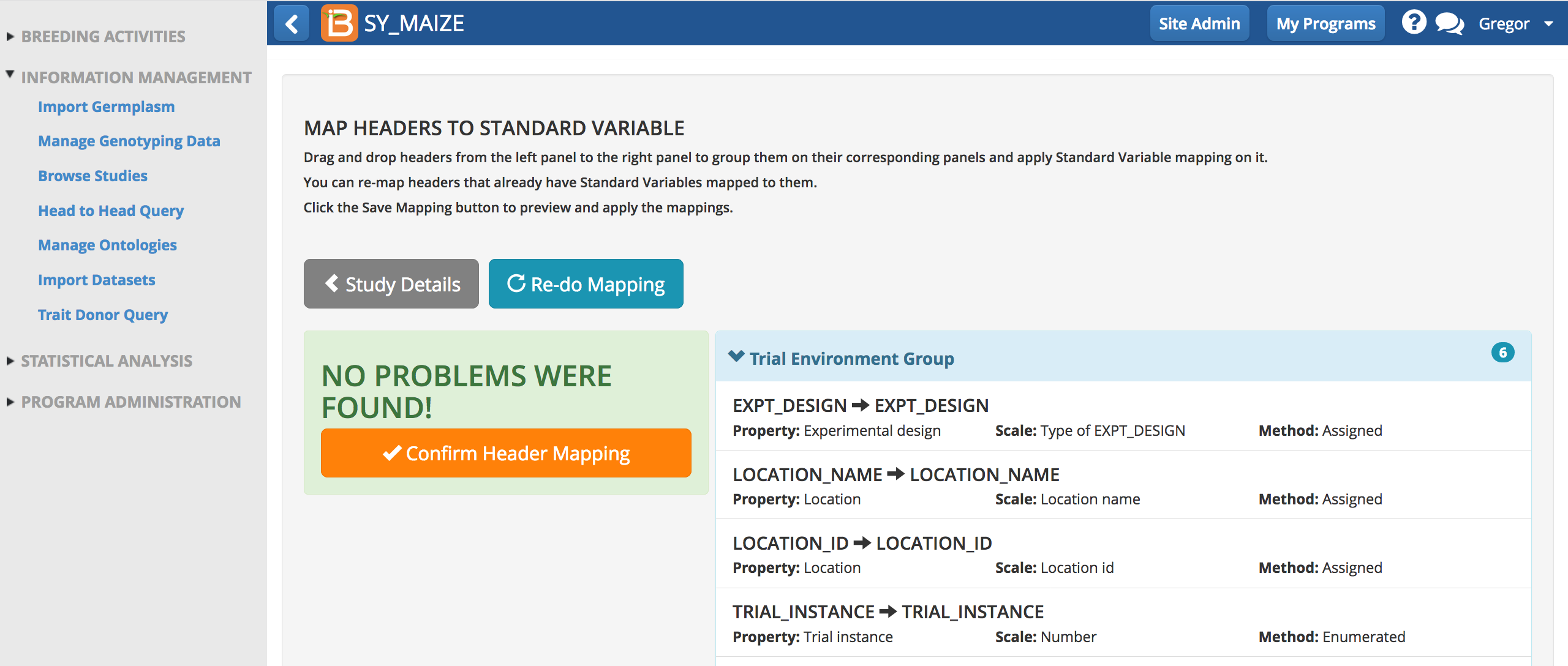
Import
- Import observations.
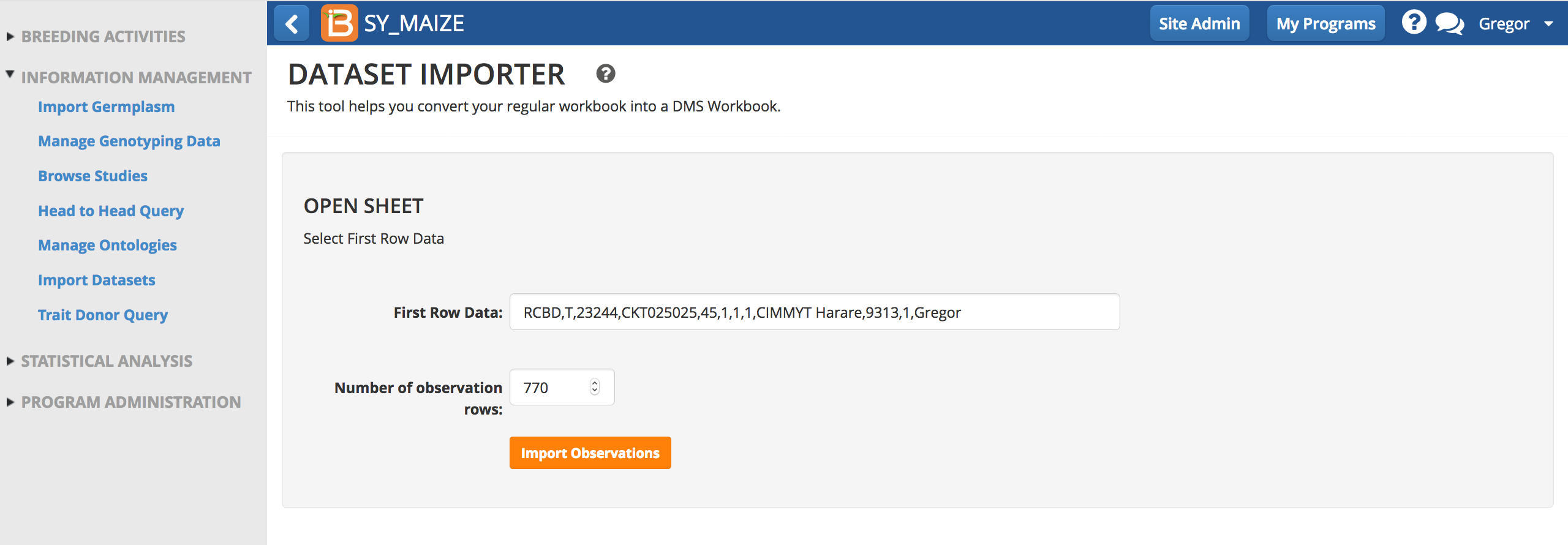
- Keep or discard out of range values (defined by the ontology).
The study is now available for analysis and queries
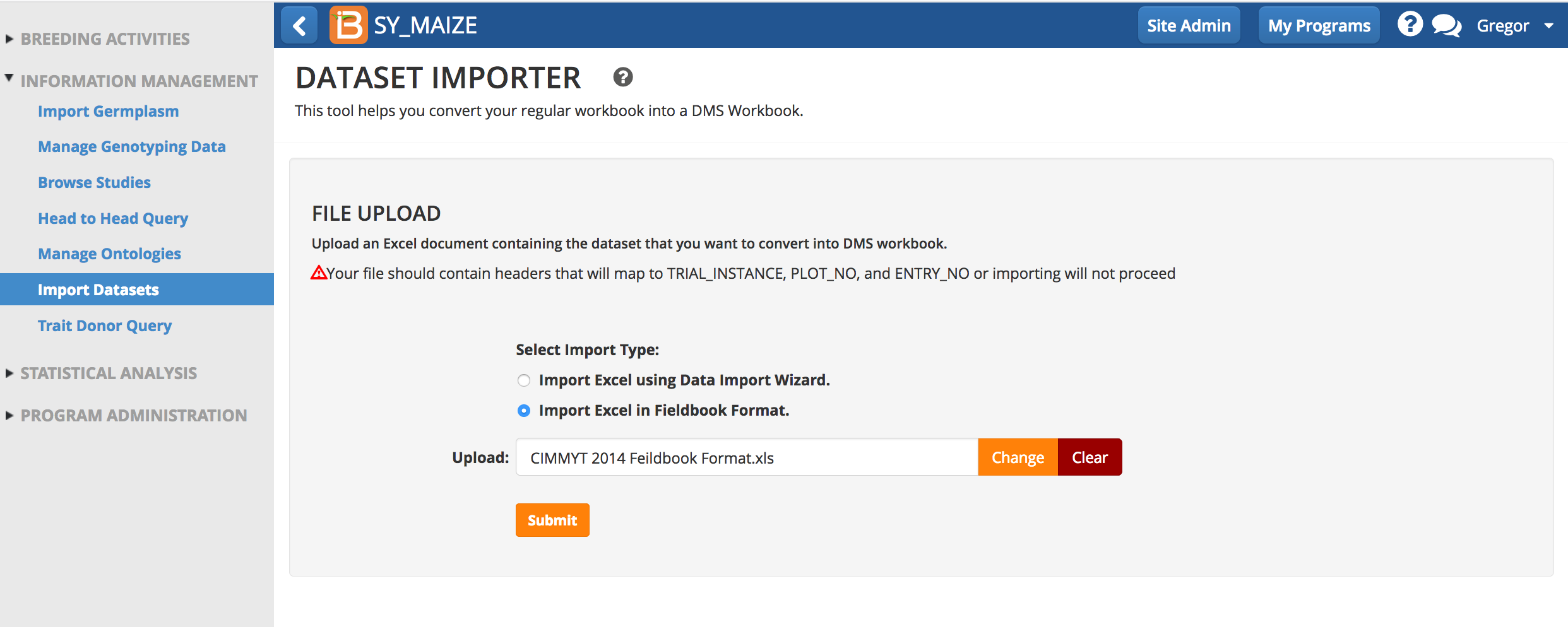
Import Fieldbook Format (beta)
Fieldbook format is now (v10) study book format. If you are currently use this feature, nothing has changed in v10 (for more information see ImportHistoricalTrialData.pdf). Expect changes and expanded documentation in the upcoming release of v11.
Related
Import Germplasm
Manage Locations
Access & User Management
Manage Studies